Comparative Genomics and Phylogenomic Analysis of the Genus Salinivibrio
- PMID: 31572321
- PMCID: PMC6749099
- DOI: 10.3389/fmicb.2019.02104
Comparative Genomics and Phylogenomic Analysis of the Genus Salinivibrio
Abstract
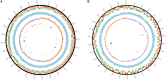
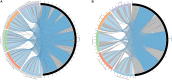
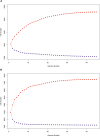
In the genomic era phylogenetic relationship among prokaryotes can be inferred from the core orthologous genes (OGs) or proteins in order to elucidate their evolutionary history and current taxonomy should benefits of that. The genus Salinivibrio belongs to the family Vibrionaceae and currently includes only five halophilic species, in spite the fact that new strains are very frequently isolated from hypersaline environments. Species belonging to this genus have undergone several reclassifications and, moreover, there are many strains of Salinivibrio with available genomes which have not been affiliated to the existing species or have been wrongly designated. Therefore, a phylogenetic study using the available genomic information is necessary to clarify the relationships of existing strains within this genus and to review their taxonomic affiliation. For that purpose, we have also sequenced the first complete genome of a Salinivibrio species, Salinivibrio kushneri AL184T, which was employed as a reference to order the contigs of the draft genomes of the type strains of the current species of this genus, as well as to perform a comparative analysis with all the other available Salinivibrio sp. genomes. The genome of S. kushneri AL184T was assembled in two circular chromosomes (with sizes of 2.84 Mb and 0.60 Mb, respectively), as typically occurs in members of the family Vibrionaceae, with nine complete ribosomal operons, which might explain the fast growing rate of salinivibrios cultured under laboratory conditions. Synteny analysis among the type strains of the genus revealed a high level of genomic conservation in both chromosomes, which allow us to hypothesize a slow speciation process or homogenization events taking place in this group of microorganisms to be tested experimentally in the future. Phylogenomic and orthologous average nucleotide identity (OrthoANI)/average amino acid identity (AAI) analyses also evidenced the elevated level of genetic relatedness within members of this genus and allowed to group all the Salinivibrio strains with available genomes in seven separated species. Genome-scale attribute study of the salinivibrios identified traits related to polar flagellum, facultatively anaerobic growth and osmotic response, in accordance to the phenotypic features described for species of this genus.
Keywords: Salinivibrio; Salinivibrio kushneri; complete genome; genomics; halophilic bacteria; hypersaline environments; phylogenomics; synteny.
Copyright © 2019 de la Haba, López-Hermoso, Sánchez-Porro, Konstantinidis and Ventosa.
Figures



Similar articles
-
Salinivibrio kushneri sp. nov., a moderately halophilic bacterium isolated from salterns.Syst Appl Microbiol. 2018 May;41(3):159-166. doi: 10.1016/j.syapm.2017.12.001. Epub 2017 Dec 21. Syst Appl Microbiol. 2018. PMID: 29331569
-
Characterization of Salinivibrio socompensis sp. nov., A New Halophilic Bacterium Isolated from the High-Altitude Hypersaline Lake Socompa, Argentina.Microorganisms. 2019 Aug 5;7(8):241. doi: 10.3390/microorganisms7080241. Microorganisms. 2019. PMID: 31387286 Free PMC article.
-
Emended description of Salinivibrio proteolyticus, including Salinivibrio costicola subsp. vallismortis and five new isolates.Int J Syst Evol Microbiol. 2018 May;68(5):1599-1607. doi: 10.1099/ijsem.0.002716. Epub 2018 Mar 26. Int J Syst Evol Microbiol. 2018. PMID: 29580324
-
Assessment of MultiLocus Sequence Analysis As a Valuable Tool for the Classification of the Genus Salinivibrio.Front Microbiol. 2017 Jun 22;8:1107. doi: 10.3389/fmicb.2017.01107. eCollection 2017. Front Microbiol. 2017. PMID: 28690592 Free PMC article.
-
Genomic and phenotypic attributes of novel salinivibrios from stromatolites, sediment and water from a high altitude lake.BMC Genomics. 2014 Jun 13;15:473. doi: 10.1186/1471-2164-15-473. BMC Genomics. 2014. PMID: 24927949 Free PMC article.
Cited by
-
Draft Genome Sequence of Saccharomonospora piscinae KCTC 19743T, an Actinobacterium Containing Secondary Metabolite Biosynthetic Gene Clusters.Microbiol Resour Announc. 2020 Apr 9;9(15):e01588-19. doi: 10.1128/MRA.01588-19. Microbiol Resour Announc. 2020. PMID: 32273373 Free PMC article.
-
Accumulation of Ectoines By Halophilic Bacteria Isolated from Fermented Shrimp Paste: An Adaptation Mechanism to Salinity, Temperature, and pH Stress.Curr Microbiol. 2021 Jun;78(6):2355-2366. doi: 10.1007/s00284-021-02481-1. Epub 2021 Apr 8. Curr Microbiol. 2021. PMID: 33830319
-
Taxogenomic and Comparative Genomic Analysis of the Genus Saccharomonospora Focused on the Identification of Biosynthetic Clusters PKS and NRPS.Front Microbiol. 2021 Mar 11;12:603791. doi: 10.3389/fmicb.2021.603791. eCollection 2021. Front Microbiol. 2021. PMID: 33776952 Free PMC article.
-
Genomic study and lipidomic bioassay of Leeuwenhoekiella parthenopeia: A novel rare biosphere marine bacterium that inhibits tumor cell viability.Front Microbiol. 2023 Jan 6;13:1090197. doi: 10.3389/fmicb.2022.1090197. eCollection 2022. Front Microbiol. 2023. PMID: 36687661 Free PMC article.
-
A halophilic metalloprotease from Salinivibrio sp. YH4 and its application in antioxidant peptide production.Front Microbiol. 2025 May 19;16:1595109. doi: 10.3389/fmicb.2025.1595109. eCollection 2025. Front Microbiol. 2025. PMID: 40458704 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

