Stop codons and the +4 nucleotide may influence the efficiency of G418 in rescuing nonsense mutations of the HERG gene
- PMID: 31573043
- PMCID: PMC6844634
- DOI: 10.3892/ijmm.2019.4360
Stop codons and the +4 nucleotide may influence the efficiency of G418 in rescuing nonsense mutations of the HERG gene
Abstract
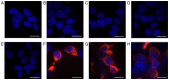
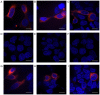
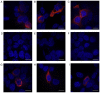
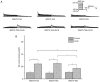
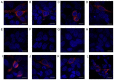
The importance of the local sequence context in determining how efficiently aminoglycosides rescue nonsense mutations has been established previously in disease models. Different stop codons appear to facilitate the termination process with differing efficiencies. Furthermore, the efficiency with which termination is suppressed may also be influenced by the local sequence context surrounding the stop codon. The strongest bias has usually been identified with the nucleotide base that immediately follows the stop codon in the majority of experiments. However, how the sequence context influences the efficiency of aminoglycosides in rescuing the human ether‑a‑go‑go‑related (HERG) protein in mammalian cells remains to be fully elucidated. Therefore, the present study was devised to examine the susceptibility of different termination codons on the HERG gene and the +4 nucleotide immediately following them to be suppressed by aminoglycosides in 293 cells. The 293 cells were transiently transfected with the wild‑type or mutant genes. The read‑through effect was subsequently examined by adding aminoglycoside G418 into the culture medium, followed by incubation of the cells for 24 h. An immunofluorescence method was then used to observe the protein expression of HERG prior to and following drug treatment. Patch clamping was performed to evaluate the function of the HERG protein. These experiments revealed that stop codons TGA and TAA in the R1014X mutant were more susceptible to treatment with the drug G418. Similar results were observed with the W927X‑TGA and W927X‑TAA mutants. Subsequently, R1014X‑TGAC, R1014X‑TGAG and R1014X‑TGAA mutants were constructed based on the R1014X‑TGAT mutant. The level of red fluorescence was observed prior to and following the administration of G418 using antibodies targeting the N‑ or C‑terminus of the HERG protein. However, the tail current density was found only to increase with the R1014X‑TGAT mutant following G418 treatment. Taken together, the results of the present study suggest that the type of premature stop codon and the context of the nucleotide immediately following at the +4 position, may determine the pharmacological rescue efficiency of the HERG gene.
Figures




Similar articles
-
Comparison of read-through effects of aminoglycosides and PTC124 on rescuing nonsense mutations of HERG gene associated with long QT syndrome.Int J Mol Med. 2014 Mar;33(3):729-35. doi: 10.3892/ijmm.2013.1601. Epub 2013 Dec 23. Int J Mol Med. 2014. PMID: 24366185
-
Stop codon context influences genome-wide stimulation of termination codon readthrough by aminoglycosides.Elife. 2020 Jan 23;9:e52611. doi: 10.7554/eLife.52611. Elife. 2020. PMID: 31971508 Free PMC article.
-
Sequence specificity of aminoglycoside-induced stop condon readthrough: potential implications for treatment of Duchenne muscular dystrophy.Ann Neurol. 2000 Aug;48(2):164-9. Ann Neurol. 2000. PMID: 10939566
-
Cancer syndromes and therapy by stop-codon readthrough.Trends Mol Med. 2012 Nov;18(11):667-78. doi: 10.1016/j.molmed.2012.09.004. Epub 2012 Oct 6. Trends Mol Med. 2012. PMID: 23044248 Review.
-
Molecular Insights into Determinants of Translational Readthrough and Implications for Nonsense Suppression Approaches.Int J Mol Sci. 2020 Dec 11;21(24):9449. doi: 10.3390/ijms21249449. Int J Mol Sci. 2020. PMID: 33322589 Free PMC article. Review.
Cited by
-
Pseudouridine-mediated stop codon readthrough in S. cerevisiae is sequence context-independent.RNA. 2020 Sep;26(9):1247-1256. doi: 10.1261/rna.076042.120. Epub 2020 May 20. RNA. 2020. PMID: 32434780 Free PMC article.
-
A system of reporters for comparative investigation of EJC-independent and EJC-enhanced nonsense-mediated mRNA decay.Nucleic Acids Res. 2024 Apr 12;52(6):e34. doi: 10.1093/nar/gkae121. Nucleic Acids Res. 2024. PMID: 38375914 Free PMC article.
References
-
- Kanters JK, Yuan L, Hedley PL, Stoevring B, Jons C, Bloch Thomsen PE, Grunnet M, Christiansen M, Jespersen T. Flecainide provocation reveals concealed brugada syndrome in a long QT syndrome family with a novel L1786Q mutation in SCN5A. Circ J. 2014;78:1136–1143. doi: 10.1253/circj.CJ-13-1167. - DOI - PubMed
-
- Marsman RF, Barc J, Beekman L, Alders M, Dooijes D, van den Wijngaard A, Ratbi I, Sefiani A, Bhuiyan ZA, Wilde AA, Bezzina CR. A mutation in CALM1 encoding calmodulin in familial idiopathic ventricular fibrillation in childhood and adolescence. J Am Coll Cardiol. 2014;63:259–266. doi: 10.1016/j.jacc.2013.07.091. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

