Compromised function of the ESCRT pathway promotes endolysosomal escape of tau seeds and propagation of tau aggregation
- PMID: 31578281
- PMCID: PMC6916486
- DOI: 10.1074/jbc.RA119.009432
Compromised function of the ESCRT pathway promotes endolysosomal escape of tau seeds and propagation of tau aggregation
Abstract
Intercellular propagation of protein aggregation is emerging as a key mechanism in the progression of several neurodegenerative diseases, including Alzheimer's disease and frontotemporal dementia (FTD). However, we lack a systematic understanding of the cellular pathways controlling prion-like propagation of aggregation. To uncover such pathways, here we performed CRISPR interference (CRISPRi) screens in a human cell-based model of propagation of tau aggregation monitored by FRET. Our screens uncovered that knockdown of several components of the endosomal sorting complexes required for transport (ESCRT) machinery, including charged multivesicular body protein 6 (CHMP6), or CHMP2A in combination with CHMP2B (whose gene is linked to familial FTD), promote propagation of tau aggregation. We found that knocking down the genes encoding these proteins also causes damage to endolysosomal membranes, consistent with a role for the ESCRT pathway in endolysosomal membrane repair. Leakiness of the endolysosomal compartment significantly enhanced prion-like propagation of tau aggregation, likely by making tau seeds more available to pools of cytoplasmic tau. Together, these findings suggest that endolysosomal escape is a critical step in tau propagation in neurodegenerative diseases.
Keywords: CRISPR/Cas; Membrane damage; Tau protein (Tau); endosomal sorting complexes required for transport (ESCRT); endosome; fluorescence resonance energy transfer (FRET); functional genomics; lysosome; protein aggregation.
© 2019 Chen et al.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures


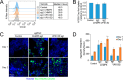
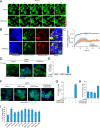

References
-
- Bejanin A., Schonhaut D. R., La Joie R., Kramer J. H., Baker S. L., Sosa N., Ayakta N., Cantwell A., Janabi M., Lauriola M., O'Neil J. P., Gorno-Tempini M. L., Miller Z. A., Rosen H. J., Miller B. L., Jagust W. J., and Rabinovici G. D. (2017) Tau pathology and neurodegeneration contribute to cognitive impairment in Alzheimer's disease. Brain 140, 3286–3300 10.1093/brain/awx243 - DOI - PMC - PubMed
-
- Berg L., McKeel D. W. Jr, Miller J. P., Storandt M., Rubin E. H., Morris J. C., Baty J., Coats M., Norton J., Goate A. M., Price J. L., Gearing M., Mirra S. S., and Saunders A. M. (1998) Clinicopathologic studies in cognitively healthy aging and Alzheimer's disease: relation of histologic markers to dementia severity, age, sex, and apolipoprotein E genotype. Arch. Neurol. 55, 326–335 10.1001/archneur.55.3.326 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- S10 OD021741/OD/NIH HHS/United States
- P50 AG023501/AG/NIA NIH HHS/United States
- K24 AG053435/AG/NIA NIH HHS/United States
- R01 NS059690/NS/NINDS NIH HHS/United States
- R56 AG057528/AG/NIA NIH HHS/United States
- P30 CA082103/CA/NCI NIH HHS/United States
- P01 AG019724/AG/NIA NIH HHS/United States
- DP2 GM119139/GM/NIGMS NIH HHS/United States
- T32 GM008568/GM/NIGMS NIH HHS/United States
- T32 GM064337/GM/NIGMS NIH HHS/United States
- U54 NS100717/NS/NINDS NIH HHS/United States
- R01 AG062359/AG/NIA NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

