Downregulation of long noncoding RNA LINC01419 inhibits cell migration, invasion, and tumor growth and promotes autophagy via inactivation of the PI3K/Akt1/mTOR pathway in gastric cancer
- PMID: 31579114
- PMCID: PMC6759708
- DOI: 10.1177/1758835919874651
Downregulation of long noncoding RNA LINC01419 inhibits cell migration, invasion, and tumor growth and promotes autophagy via inactivation of the PI3K/Akt1/mTOR pathway in gastric cancer
Retraction in
-
Retraction notice: Downregulation of long noncoding RNA LINC01419 inhibits cell migration, invasion, and tumor growth and promotes autophagy via inactivation of the PI3K/Akt1/mTOR pathway in gastric cancer.Ther Adv Med Oncol. 2021 Dec 15;13:17588359211061903. doi: 10.1177/17588359211061903. eCollection 2021. Ther Adv Med Oncol. 2021. PMID: 35035532 Free PMC article.
Abstract
Background: Accumulating evidence has highlighted the crucial role of long noncoding RNAs (lncRNAs) in the tumorigenesis of gastric cancer (GC), which is the most common gastrointestinal malignancy. The present study aimed to identify the capacity of lncRNA LINC01419 (LINC01419) in GC progression, with the potential mechanism explored.
Methods: Highly expressed lncRNAs were identified by in silico analysis, with the LINC01419 expression in GC tissues measured using reverse transcription-quantitative PCR (RT-qPCR). The GC cells were subsequently transfected with siRNA against LINC01419 or Rapamycin (the inhibitor of the mTOR pathway), or both, in order to measure cell migration and invasion in vitro as well as tumor growth and metastasis in vivo. Moreover, the expression of PI3K/Akt1/mTOR pathway-associated factors was determined.
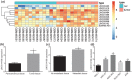
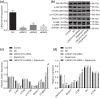
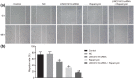
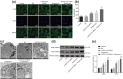
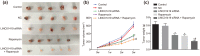
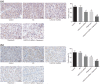
Results: LINC01419, highly expressed in GC samples of the Gene Expression Omnibus database, was observed to be markedly upregulated in GC tissues. Moreover, LINC01419 silencing, or PI3K/Akt1/mTOR pathway inhibition, exhibited an inhibitory role in GC cell migration and invasion in vitro, coupled with promoted cell autophagy in vitro, and inhibited tumor growth and metastasis in vivo. It was also revealed that LINC01419 silencing blocked the PI3K/Akt1/mTOR pathway, as proved by decreased extents of Akt1 and mTOR phosphorylation.
Conclusions: In conclusion, LINC01419 inhibition may suppress GC cell invasion and migration, and promote autophagy via inhibition of the PI3K/Akt1/mTOR pathway. This provides significant theoretical basis and possibilities for further elucidation of the molecular mechanism of GC and finding new molecular-targeted therapeutic regimens.
Keywords: LINC01419; RNA; autophagy; gastric cancer; invasion and migration; long noncoding PI3K/Akt1/mTOR pathway.
© The Author(s), 2019.
Conflict of interest statement
Conflict of interest statement: The authors declare no conflicts of interest in preparing this article.
Figures

Similar articles
-
Retraction notice: Downregulation of long noncoding RNA LINC01419 inhibits cell migration, invasion, and tumor growth and promotes autophagy via inactivation of the PI3K/Akt1/mTOR pathway in gastric cancer.Ther Adv Med Oncol. 2021 Dec 15;13:17588359211061903. doi: 10.1177/17588359211061903. eCollection 2021. Ther Adv Med Oncol. 2021. PMID: 35035532 Free PMC article.
-
STAT3-induced lncRNA HAGLROS overexpression contributes to the malignant progression of gastric cancer cells via mTOR signal-mediated inhibition of autophagy.Mol Cancer. 2018 Jan 12;17(1):6. doi: 10.1186/s12943-017-0756-y. Mol Cancer. 2018. PMID: 29329543 Free PMC article.
-
LINC01419 promotes cell proliferation and metastasis in hepatocellular carcinoma by enhancing NDRG1 promoter activity.Cell Oncol (Dordr). 2020 Oct;43(5):931-947. doi: 10.1007/s13402-020-00540-6. Epub 2020 Jun 15. Cell Oncol (Dordr). 2020. PMID: 32557341
-
Potential mechanism of traditional Chinese medicine intervention in gastric cancer: targeted regulation of autophagy.Front Pharmacol. 2025 Feb 18;16:1548672. doi: 10.3389/fphar.2025.1548672. eCollection 2025. Front Pharmacol. 2025. PMID: 40066330 Free PMC article. Review.
-
Autophagy Related Noncoding RNAs: Emerging Regulatory Factors of Gastric Cancer.Cancer Manag Res. 2022 Jul 19;14:2215-2224. doi: 10.2147/CMAR.S364761. eCollection 2022. Cancer Manag Res. 2022. PMID: 35898946 Free PMC article. Review.
Cited by
-
Comprehensive Analysis of Ferroptosis-Related LncRNAs in Breast Cancer Patients Reveals Prognostic Value and Relationship With Tumor Immune Microenvironment.Front Surg. 2021 Oct 4;8:742360. doi: 10.3389/fsurg.2021.742360. eCollection 2021. Front Surg. 2021. PMID: 34671639 Free PMC article.
-
Fatty Acid-Binding Protein 4 (FABP4) Suppresses Proliferation and Migration of Endometrial Cancer Cells via PI3K/Akt Pathway.Onco Targets Ther. 2021 Jun 29;14:3929-3942. doi: 10.2147/OTT.S311792. eCollection 2021. Onco Targets Ther. 2021. PMID: 34234461 Free PMC article.
-
The lncRNAs MIR600HG and TSPOAP1-AS1 may potentially act as biomarkers for predicting pancreatic cancer.Transl Cancer Res. 2020 Feb;9(2):809-817. doi: 10.21037/tcr.2019.12.09. Transl Cancer Res. 2020. PMID: 35117426 Free PMC article.
-
A Novel Autophagy-Related Long Non-Coding RNA Signature to Predict Prognosis and Therapeutic Response in Esophageal Squamous Cell Carcinoma.Int J Gen Med. 2021 Nov 16;14:8325-8339. doi: 10.2147/IJGM.S333697. eCollection 2021. Int J Gen Med. 2021. PMID: 34815705 Free PMC article.
-
Forkhead box K1 regulates the malignant behavior of gastric cancer by inhibiting autophagy.Ann Transl Med. 2020 Feb;8(4):107. doi: 10.21037/atm.2019.12.123. Ann Transl Med. 2020. PMID: 32175400 Free PMC article.
References
Publication types
LinkOut - more resources
Full Text Sources
Miscellaneous

