Integrated molecular characterization of chondrosarcoma reveals critical determinants of disease progression
- PMID: 31604924
- PMCID: PMC6789144
- DOI: 10.1038/s41467-019-12525-7
Integrated molecular characterization of chondrosarcoma reveals critical determinants of disease progression
Abstract
Chondrosarcomas are primary cancers of cartilaginous tissue with highly contrasting prognoses. These tumors are defined by recurrent mutations in the IDH genes and other genetic alterations including inactivation of CDKN2A and COL2A1; however, these have no clinical value. Here we use multi-omics molecular profiles from a series of cartilage tumors and find an mRNA classification that identifies two subtypes of chondrosarcomas defined by a balance in tumor differentiation and cell cycle activation. The microRNA classification reveals the importance of the loss of expression of the 14q32 locus in defining the level of malignancy. Finally, DNA methylation is associated with IDH mutations. We can use the multi-omics classifications to predict outcome. We propose an mRNA-only classifier to reproduce the integrated multi-omics classification, and its application to relapsed tumor samples shows the progressive nature of the classification. Thus, it may be possible to use mRNA-based signatures to detect patients with high-risk chondrosarcomas.
Conflict of interest statement
The authors declare no competing interests.
Figures


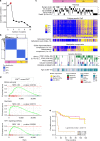

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

