Analysis of 1,000 Type-Strain Genomes Improves Taxonomic Classification of Bacteroidetes
- PMID: 31608019
- PMCID: PMC6767994
- DOI: 10.3389/fmicb.2019.02083
Analysis of 1,000 Type-Strain Genomes Improves Taxonomic Classification of Bacteroidetes
Abstract
Although considerable progress has been made in recent years regarding the classification of bacteria assigned to the phylum Bacteroidetes, there remains a need to further clarify taxonomic relationships within a diverse assemblage that includes organisms of clinical, piscicultural, and ecological importance. Bacteroidetes classification has proved to be difficult, not least when taxonomic decisions rested heavily on interpretation of poorly resolved 16S rRNA gene trees and a limited number of phenotypic features. Here, draft genome sequences of a greatly enlarged collection of genomes of more than 1,000 Bacteroidetes and outgroup type strains were used to infer phylogenetic trees from genome-scale data using the principles drawn from phylogenetic systematics. The majority of taxa were found to be monophyletic but several orders, families and genera, including taxa proposed long ago such as Bacteroides, Cytophaga, and Flavobacterium but also quite recent taxa, as well as a few species were shown to be in need of revision. According proposals are made for the recognition of new orders, families and genera, as well as the transfer of a variety of species to other genera. In addition, emended descriptions are given for many species mainly involving information on DNA G+C content and (approximate) genome size, both of which can be considered valuable taxonomic markers. We detected many incongruities when comparing the results of the present study with existing classifications, which appear to be caused by insufficiently resolved 16S rRNA gene trees or incomplete taxon sampling. The few significant incongruities found between 16S rRNA gene and whole genome trees underline the pitfalls inherent in phylogenies based upon single gene sequences and the impediment in using ordinary bootstrapping in phylogenomic studies, particularly when combined with too narrow gene selections. While a significant degree of phylogenetic conservation was detected in all phenotypic characters investigated, the overall fit to the tree varied considerably, which is one of the probable causes of misclassifications in the past, much like the use of plesiomorphic character states as diagnostic features.
Keywords: G+C content; chemotaxonomy; genome BLAST distance phylogeny; genome size; morphology; phylogenetic systematics; phylogenomics.
Copyright © 2019 García-López, Meier-Kolthoff, Tindall, Gronow, Woyke, Kyrpides, Hahnke and Göker.
Figures

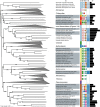






References
-
- Albert R. A., Zitomer D., Dollhopf M., Schauer-Gimenez A. E., Struble C., King M., et al. (2014). Proposal of Vibrionimonas magnilacihabitans gen. nov., sp. nov., a curved Gram-stain-negative bacterium isolated from lake water. Int. J. Syst. Evol. Microbiol. 64, 613–620. 10.1099/ijs.0.056663-0 - DOI - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

