Understanding the genetics of neuropsychiatric disorders: the potential role of genomic regulatory blocks
- PMID: 31616042
- PMCID: PMC6906185
- DOI: 10.1038/s41380-019-0518-x
Understanding the genetics of neuropsychiatric disorders: the potential role of genomic regulatory blocks
Abstract
Recent genome-wide association studies have identified numerous loci associated with neuropsychiatric disorders. The majority of these are in non-coding regions, and are commonly assigned to the nearest gene along the genome. However, this approach neglects the three-dimensional organisation of the genome, and the fact that the genome contains arrays of extremely conserved non-coding elements termed genomic regulatory blocks (GRBs), which can be utilized to detect genes under long-range developmental regulation. Here we review a GRB-based approach to assign loci in non-coding regions to potential target genes, and apply it to reanalyse the results of one of the largest schizophrenia GWAS (SWG PGC, 2014). We further apply this approach to GWAS data from two related neuropsychiatric disorders-autism spectrum disorder and bipolar disorder-to show that it is applicable to developmental disorders in general. We find that disease-associated SNPs are overrepresented in GRBs and that the GRB model is a powerful tool for linking these SNPs to their correct target genes under long-range regulation. Our analysis identifies novel genes not previously implicated in schizophrenia and corroborates a number of predicted targets from the original study. The results are available as an online resource in which the genomic context and the strength of enhancer-promoter associations can be browsed for each schizophrenia-associated SNP.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures
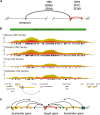


References
-
- Bechtel W. Mechanists must be holists too! perspectives from circadian biology William. J Hist Biol. 2016;49:705–31. - PubMed
-
- Bechtel W. Network Organization in Health and Disease: on being a reductionist and a systems biologist too. Pharmacopsychiatry. 2013;46:10–21. - PubMed
-
- Reich DE, Cargill M, Bolk S, Ireland J, Sabeti PC, Richter DJ, et al. Link disequilibrium in the human genome. 2001;9:199–204. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

