Nanosecond photochemically promoted click chemistry for enhanced neuropeptide visualization and rapid protein labeling
- PMID: 31619683
- PMCID: PMC6795811
- DOI: 10.1038/s41467-019-12548-0
Nanosecond photochemically promoted click chemistry for enhanced neuropeptide visualization and rapid protein labeling
Abstract
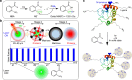
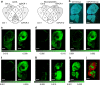
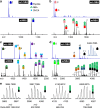
Comprehensive protein identification and concomitant structural probing of proteins are of great biological significance. However, this is challenging to accomplish simultaneously in one confined space. Here, we develop a nanosecond photochemical reaction (nsPCR)-based click chemistry, capable of structural probing of proteins and enhancing their identifications through on-demand removal of surrounding matrices within nanoseconds. The nsPCR is initiated using a photoactive compound, 2-nitrobenzaldehyde (NBA), and is examined by matrix-assisted laser desorption/ionization-mass spectrometry (MALDI-MS). Benefiting from the on-demand matrix-removal effect, this nsPCR strategy enables enhanced neuropeptide identification and visualization from complex tissue samples such as mouse brain tissue. The design shows great promise for structural probing of proteins up to 155 kDa due to the exclusive accessibility of nsPCR to primary amine groups, as demonstrated by its general applicability using a series of proteins with various lysine residues from multiple sample sources, with accumulated labeling efficiencies greater than 90%.
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Nanosecond Photochemical Reaction for Enhanced Identification, Quantification, and Visualization of Primary Amine-Containing Metabolites by MALDI-Mass Spectrometry.Anal Chem. 2022 Mar 8;94(9):3774-3781. doi: 10.1021/acs.analchem.1c03840. Epub 2022 Feb 21. Anal Chem. 2022. PMID: 35189681 Free PMC article.
-
Nanosecond Photochemical Reaction (nsPCR) for Enhanced Mass Spectrometric Identification, Quantification, and Visualization of Metabolites and Neuropeptides.Methods Mol Biol. 2022;2437:143-157. doi: 10.1007/978-1-0716-2030-4_10. Methods Mol Biol. 2022. PMID: 34902146 Free PMC article.
-
A Simple and Effective Sample Preparation Strategy for MALDI-MS Imaging of Neuropeptide Changes in the Crustacean Brain Due to Hypoxia and Hypercapnia Stress.J Am Soc Mass Spectrom. 2020 May 6;31(5):1058-1065. doi: 10.1021/jasms.9b00107. Epub 2020 Mar 23. J Am Soc Mass Spectrom. 2020. PMID: 32150406 Free PMC article.
-
Surface-assisted laser desorption ionization mass spectrometry techniques for application in forensics.Mass Spectrom Rev. 2015 Nov-Dec;34(6):627-40. doi: 10.1002/mas.21431. Epub 2014 Jun 11. Mass Spectrom Rev. 2015. PMID: 24916100 Review.
-
Discovering new invertebrate neuropeptides using mass spectrometry.Mass Spectrom Rev. 2006 Jan-Feb;25(1):77-98. doi: 10.1002/mas.20055. Mass Spectrom Rev. 2006. PMID: 15937922 Review.
Cited by
-
Mass spectrometry imaging: the rise of spatially resolved single-cell omics.Nat Methods. 2023 Mar;20(3):327-330. doi: 10.1038/s41592-023-01774-6. Nat Methods. 2023. PMID: 36899158 No abstract available.
-
Insight to Functional Conformation and Noncovalent Interactions of Protein-Protein Assembly Using MALDI Mass Spectrometry.Molecules. 2020 Oct 28;25(21):4979. doi: 10.3390/molecules25214979. Molecules. 2020. PMID: 33126406 Free PMC article. Review.
-
Neuropeptidomics: Improvements in Mass Spectrometry Imaging Analysis and Recent Advancements.Curr Protein Pept Sci. 2021;22(2):158-169. doi: 10.2174/1389203721666201116115708. Curr Protein Pept Sci. 2021. PMID: 33200705 Free PMC article. Review.
-
Nanosecond Photochemical Reaction for Enhanced Identification, Quantification, and Visualization of Primary Amine-Containing Metabolites by MALDI-Mass Spectrometry.Anal Chem. 2022 Mar 8;94(9):3774-3781. doi: 10.1021/acs.analchem.1c03840. Epub 2022 Feb 21. Anal Chem. 2022. PMID: 35189681 Free PMC article.
-
Complementary neuropeptide detection in crustacean brain by mass spectrometry imaging using formalin and alternative aqueous tissue washes.Anal Bioanal Chem. 2021 Apr;413(10):2665-2673. doi: 10.1007/s00216-020-03073-x. Epub 2021 Jan 5. Anal Bioanal Chem. 2021. PMID: 33403426 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
- RF1AG052324/U.S. Department of Health & Human Services | National Institutes of Health (NIH)/International
- R56 DK071801/DK/NIDDK NIH HHS/United States
- R56 MH110215/MH/NIMH NIH HHS/United States
- S10 RR029531/RR/NCRR NIH HHS/United States
- R56MH110215/U.S. Department of Health & Human Services | National Institutes of Health (NIH)/International
- RF1 AG052324/AG/NIA NIH HHS/United States
- R01 DK071801/DK/NIDDK NIH HHS/United States
- 1F31GM126870/U.S. Department of Health & Human Services | National Institutes of Health (NIH)/International
- U01 CA231081/CA/NCI NIH HHS/United States
- S10 OD025084/OD/NIH HHS/United States
- F31 GM126870/GM/NIGMS NIH HHS/United States
- U01CA231081/U.S. Department of Health & Human Services | National Institutes of Health (NIH)/International
LinkOut - more resources
Full Text Sources
Other Literature Sources

