Global Secretome Characterization of the Pathogenic Yeast Candida glabrata
- PMID: 31621333
- PMCID: PMC6947636
- DOI: 10.1021/acs.jproteome.9b00299
Global Secretome Characterization of the Pathogenic Yeast Candida glabrata
Abstract
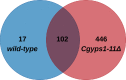
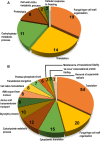
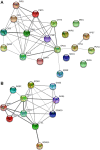
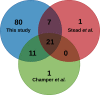
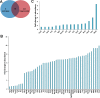
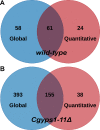
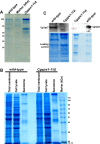
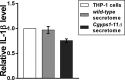
Secretory proteins are key modulators of host-pathogen interaction. The human opportunistic fungal pathogen Candida glabrata lacks secreted proteolytic activity but possesses 11 glycosylphosphatidylinositol-anchored aspartyl proteases, also referred to as Yapsins (CgYps1-11), that are essential for its virulence. To delineate the role of CgYapsins in interaction with host cells, we have profiled, through liquid chromatography-tandem mass spectrometry (LC-MS/MS) approach, the total secretome of wild-type and Cgyps1-11Δ mutant. The wild-type secretome consisted of 119 proteins which were primarily involved in cell wall organization, carbohydrate metabolism, proteolysis, and translation processes. Of eight CgYapsins identified in the secretome, the release of two major CgYapsins, CgYps1 and CgYps7, to the medium was confirmed by Western analysis. Further, comparative analysis revealed 20 common proteins, probably signifying the core fungal secretome, among C. glabrata, Saccharomyces cerevisiae, and Candida albicans secretomes. Strikingly, the Cgyps1-11Δ secretome was 4.6-fold larger, and contained 65 differentially abundant proteins, as revealed by label-free quantitative profiling, with 49 and 16 being high- and low-abundant proteins, respectively, compared to the wild-type secretome. Importantly, the CgMsb2 mucin, a putative CgYapsins' substrate, was six-fold underrepresented in the mutant secretome. Altogether, we demonstrate for the first time that CgYapsins are both bona fide constituents and key modulators of the C. glabrata secretome.
Keywords: GPI proteins; Yapsins; carbohydrate metabolism; cell wall organization; core fungal secretome; pathogenic fungi; secretory signal peptide.
Conflict of interest statement
The authors declare no competing financial interest.
Figures

Similar articles
-
The yapsin family of aspartyl proteases regulate glucose homeostasis in Candida glabrata.J Biol Chem. 2022 Feb;298(2):101593. doi: 10.1016/j.jbc.2022.101593. Epub 2022 Jan 17. J Biol Chem. 2022. PMID: 35051415 Free PMC article.
-
Aspartyl proteases in Candida glabrata are required for suppression of the host innate immune response.J Biol Chem. 2018 Apr 27;293(17):6410-6433. doi: 10.1074/jbc.M117.813741. Epub 2018 Feb 28. J Biol Chem. 2018. PMID: 29491142 Free PMC article.
-
Polymorphism in the regulatory regions of genes CgYPS1 and CgYPS7 encoding yapsins in Candida glabrata is associated with changes in expression levels.FEMS Yeast Res. 2017 Dec 1;17(8). doi: 10.1093/femsyr/fox077. FEMS Yeast Res. 2017. PMID: 29069395
-
The mating type-like loci of Candida glabrata.Rev Iberoam Micol. 2014 Jan-Mar;31(1):30-4. doi: 10.1016/j.riam.2013.09.016. Epub 2013 Nov 16. Rev Iberoam Micol. 2014. PMID: 24252826 Review.
-
Extracellular proteinases of Candida species pathogenic yeasts.Mol Oral Microbiol. 2018 Apr;33(2):113-124. doi: 10.1111/omi.12206. Epub 2018 Jan 19. Mol Oral Microbiol. 2018. PMID: 29139623 Review.
Cited by
-
The secreted acid trehalase encoded by the CgATH1 gene is involved in Candida glabrata virulence.Mem Inst Oswaldo Cruz. 2020 Oct 30;115:e200401. doi: 10.1590/0074-02760200401. eCollection 2020. Mem Inst Oswaldo Cruz. 2020. PMID: 33146242 Free PMC article.
-
The yapsin family of aspartyl proteases regulate glucose homeostasis in Candida glabrata.J Biol Chem. 2022 Feb;298(2):101593. doi: 10.1016/j.jbc.2022.101593. Epub 2022 Jan 17. J Biol Chem. 2022. PMID: 35051415 Free PMC article.
-
Aspartyl proteases target host actin nucleator complex protein to limit epithelial innate immunity.EMBO Rep. 2024 Nov;25(11):4846-4875. doi: 10.1038/s44319-024-00270-y. Epub 2024 Sep 30. EMBO Rep. 2024. PMID: 39349750 Free PMC article.
-
Characterization of the Secretome of Pathogenic Candida glabrata and Their Effectiveness against Systemic Candidiasis in BALB/c Mice for Vaccine Development.Pharmaceutics. 2022 Sep 21;14(10):1989. doi: 10.3390/pharmaceutics14101989. Pharmaceutics. 2022. PMID: 36297425 Free PMC article.
-
Candidiasis: Insights into Virulence Factors, Complement Evasion and Antifungal Drug Resistance.Microorganisms. 2025 Jan 25;13(2):272. doi: 10.3390/microorganisms13020272. Microorganisms. 2025. PMID: 40005639 Free PMC article. Review.
References
-
- Pfaller M. A.; Pappas P. G.; Wingard J. R. Invasive Fungal Pathogens: Current Epidemiological Trends. Clin. Infect. Dis. 2006, 43, S3–S14. 10.1086/504490. - DOI
-
- Colombo A. L.; Guimarães T.; Sukienik T.; Pasqualotto A. C.; Andreotti R.; Queiroz-Telles F.; Nouér S. A.; Nucci M. Prognostic Factors and Historical Trends in the Epidemiology of Candidemia in Critically Ill Patients: An Analysis of Five Multicenter Studies Sequentially Conducted over a 9-Year Period. Intensive Care Med. 2014, 40, 1489–1498. 10.1007/s00134-014-3400-y. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

