An independently validated survival nomogram for lower-grade glioma
- PMID: 31621885
- PMCID: PMC7229246
- DOI: 10.1093/neuonc/noz191
An independently validated survival nomogram for lower-grade glioma
Abstract
Background: Gliomas are the most common primary malignant brain tumor. Diffuse low-grade and intermediate-grade gliomas, which together compose the lower-grade gliomas (LGGs; World Health Organization [WHO] grades II and III), present a therapeutic challenge to physicians due to the heterogeneity of their clinical behavior. Nomograms are useful tools for individualized estimation of survival. This study aimed to develop and independently validate a survival nomogram for patients with newly diagnosed LGG.
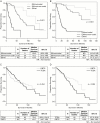
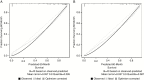
Methods: Data were obtained for newly diagnosed LGG patients from The Cancer Genome Atlas (TCGA) and the Ohio Brain Tumor Study (OBTS) with the following variables: tumor grade (II or III), age at diagnosis, sex, Karnofsky performance status (KPS), and molecular subtype (IDH mutant with 1p/19q codeletion [IDHmut-codel], IDH mutant without 1p/19q codeletion, and IDH wild-type). Survival was assessed using Cox proportional hazards regression, random survival forests, and recursive partitioning analysis, with adjustment for known prognostic factors. The models were developed using TCGA data and independently validated using the OBTS data. Models were internally validated using 10-fold cross-validation and externally validated with calibration curves.
Results: A final nomogram was validated for newly diagnosed LGG. Factors that increased the probability of survival included grade II tumor, younger age at diagnosis, having a high KPS, and the IDHmut-codel molecular subtype.
Conclusions: A nomogram that calculates individualized survival probabilities for patients with newly diagnosed LGG could be useful to health care providers for counseling patients regarding treatment decisions and optimizing therapeutic approaches. Free online software for implementing this nomogram is provided: https://hgittleman.shinyapps.io/LGG_Nomogram_H_Gittleman/.
Key points: 1. A survival nomogram for lower-grade glioma patients has been developed and externally validated.2. Free online software for implementing this nomogram is provided allowing for ease of use by practicing health care providers.
Keywords: TCGA; glioma; lower grade glioma; nomogram; survival.
© The Author(s) 2019. Published by Oxford University Press on behalf of the Society for Neuro-Oncology. All rights reserved. For permissions, please e-mail: journals.permissions@oup.com.
Figures

Comment in
-
A validated prognostic nomogram for patients with newly diagnosed lower-grade gliomas in a large-scale Asian cohort.Neuro Oncol. 2020 May 15;22(5):729-731. doi: 10.1093/neuonc/noaa027. Neuro Oncol. 2020. PMID: 32025722 Free PMC article. No abstract available.
Similar articles
-
An independently validated nomogram for isocitrate dehydrogenase-wild-type glioblastoma patient survival.Neurooncol Adv. 2019 May-Dec;1(1):vdz007. doi: 10.1093/noajnl/vdz007. Epub 2019 May 30. Neurooncol Adv. 2019. PMID: 31608326 Free PMC article.
-
The T2-FLAIR-mismatch sign as an imaging biomarker for IDH and 1p/19q status in diffuse low-grade gliomas: a systematic review with a Bayesian approach to evaluation of diagnostic test performance.Neurosurg Focus. 2019 Dec 1;47(6):E13. doi: 10.3171/2019.9.FOCUS19660. Neurosurg Focus. 2019. PMID: 31786548
-
Conventional MR-based Preoperative Nomograms for Prediction of IDH/1p19q Subtype in Low-Grade Glioma.Acad Radiol. 2019 Aug;26(8):1062-1070. doi: 10.1016/j.acra.2018.09.022. Epub 2018 Nov 2. Acad Radiol. 2019. PMID: 30393056
-
Advanced imaging parameters improve the prediction of diffuse lower-grade gliomas subtype, IDH mutant with no 1p19q codeletion: added value to the T2/FLAIR mismatch sign.Eur Radiol. 2020 Feb;30(2):844-854. doi: 10.1007/s00330-019-06395-2. Epub 2019 Aug 24. Eur Radiol. 2020. PMID: 31446467
-
Radiation and chemotherapy for high-risk lower grade gliomas: Choosing between temozolomide and PCV.Cancer Med. 2020 Jan;9(1):3-11. doi: 10.1002/cam4.2686. Epub 2019 Nov 7. Cancer Med. 2020. PMID: 31701682 Free PMC article. Review.
Cited by
-
Prognostic Prediction Model for Glioblastoma: A Metabolic Gene Signature and Independent External Validation.J Cancer. 2021 May 5;12(13):3796-3808. doi: 10.7150/jca.53827. eCollection 2021. J Cancer. 2021. PMID: 34093788 Free PMC article.
-
A novel DNA repair-related nomogram predicts survival in low-grade gliomas.CNS Neurosci Ther. 2021 Feb;27(2):186-195. doi: 10.1111/cns.13464. Epub 2020 Oct 16. CNS Neurosci Ther. 2021. PMID: 33063446 Free PMC article.
-
MACF1 mutations predict poor prognosis: a novel potential therapeutic target for breast cancer.Am J Transl Res. 2022 Nov 15;14(11):7670-7688. eCollection 2022. Am J Transl Res. 2022. PMID: 36505342 Free PMC article.
-
Expression and prognostic potential of GPX1 in human cancers based on data mining.Ann Transl Med. 2020 Feb;8(4):124. doi: 10.21037/atm.2020.02.36. Ann Transl Med. 2020. PMID: 32175417 Free PMC article.
-
Clinical features and risk factors analysis of bronchitis obliterans due to refractory Mycoplasma pneumoniae pneumonia in children: a nomogram prediction model.BMC Infect Dis. 2021 Oct 21;21(1):1085. doi: 10.1186/s12879-021-06783-4. BMC Infect Dis. 2021. PMID: 34674642 Free PMC article.
References
-
- Goodenberger ML, Jenkins RB. Genetics of adult glioma. Cancer Genet. 2012;205(12):613–621. - PubMed
-
- Gorovets D, Kannan K, Shen R, et al. . IDH mutation and neuroglial developmental features define clinically distinct subclasses of lower grade diffuse astrocytic glioma. Clin Cancer Res. 2012;18(9):2490–2501. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

