A flavin-dependent monooxgenase confers resistance to chlorantraniliprole in the diamondback moth, Plutella xylostella
- PMID: 31626952
- PMCID: PMC6880784
- DOI: 10.1016/j.ibmb.2019.103247
A flavin-dependent monooxgenase confers resistance to chlorantraniliprole in the diamondback moth, Plutella xylostella
Abstract
The diamondback moth, Plutella xylostella, is a damaging pest of cruciferous crops, and has evolved resistance to many of the insecticides used for control, including members of the diamide class. Previous work on the molecular basis of resistance to diamides has documented mutations in the target-site, the ryanodine receptor, in resistant populations of P. xylostella worldwide. In contrast the role of metabolic resistance to this insecticide class is significantly less clear. Here we show that overexpression of a flavin-dependent monooxgenase (FMO) confers resistance to the diamide chlorantraniliprole in P. xylostella. Transcriptome profiling of diamide resistant strains, with and without target-site resistance, revealed constitutive over-expression of several transcripts encoding detoxification enzymes compared to susceptible strains. Two of these, CYP6BG1, and PxFMO2 were particularly highly overexpressed (33,000 and 14,700-fold, respectively) in a resistant strain (HAW) lacking target-site resistance. After 17 generations without diamide selection the resistance of the HAW strain fell by 52-fold and the expression of PxFMO2 by > 1300-fold, however, the expression of CYP6BG1 declined by only 3-fold. Generation of transgenic Drosophila melanogaster expressing these genes demonstrated that PxFMO2, but not CYP6BG1, confers resistance in vivo. Overexpression of PxFMO2 in the HAW strain is associated with mutations, including a putative transposable element insertion, in the promoter of this gene. These enhance the expression of a reporter gene when expressed in a lepidopteran cell line suggesting they are, at least in part, responsible for the overexpression of PxFMO2 in the resistant strain. Our results provide new evidence that insect FMOs can be recruited to provide resistance to synthetic insecticides.
Keywords: Chlorantraniliprole; Diamide; Flavin monooxygenase; Insecticides; Plutella xylostella; Resistance.
Copyright © 2019 The Author(s). Published by Elsevier Ltd.. All rights reserved.
Figures
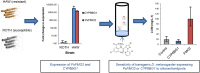




References
-
- Abbott W.S. A method for computing the effectiveness of an insecticide. J. Econ. Entomol. 1925;18:265–267.
-
- Bautista M.A.M., Miyata T., Miura K., Tanaka T. RNA interference-mediated knockdown of a cytochrome P450, CYP6BG1, from the diamondback moth, Plutella xylostella, reduces larval resistance to permethrin. Insect Biochem. Mol. Biol. 2009;39:38–46. - PubMed
-
- Douris V., Papapostolou K.M., Ilias A., Roditakis E., Kounadi S., Riga M., Nauen R., Vontas J. Investigation of the contribution of RyR target-site mutations in diamide resistance by CRISPR/Cas9 genome modification in Drosophila. Insect Biochem. Mol. Biol. 2017;87:127–135. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

