Genera of phytopathogenic fungi: GOPHY 3
- PMID: 31636728
- PMCID: PMC6797016
- DOI: 10.1016/j.simyco.2019.05.001
Genera of phytopathogenic fungi: GOPHY 3
Abstract
This paper represents the third contribution in the Genera of Phytopathogenic Fungi (GOPHY) series. The series provides morphological descriptions, information about the pathology, distribution, hosts and disease symptoms for the treated genera, as well as primary and secondary DNA barcodes for the currently accepted species included in these. This third paper in the GOPHY series treats 21 genera of phytopathogenic fungi and their relatives including: Allophoma, Alternaria, Brunneosphaerella, Elsinoe, Exserohilum, Neosetophoma, Neostagonospora, Nothophoma, Parastagonospora, Phaeosphaeriopsis, Pleiocarpon, Pyrenophora, Ramichloridium, Seifertia, Seiridium, Septoriella, Setophoma, Stagonosporopsis, Stemphylium, Tubakia and Zasmidium. This study includes three new genera, 42 new species, 23 new combinations, four new names, and three typifications of older names.
Keywords: Allophoma pterospermicola Q. Chen & L. Cai; Alternaria aconidiophora Iturrieta-González, Dania García & Gené; Alternaria altcampina Iturrieta-González, Dania García & Gené; Alternaria chlamydosporifera Iturrieta-González, Dania García & Gené; Alternaria curvata Iturrieta-González, Dania García & Gené; Alternaria fimeti Iturrieta-González, Dania García & Gené; Alternaria inflata Iturrieta-González, Dania García & Gené; Alternaria lawrencei Iturrieta-González, Dania García & Gené; Alternaria montsantina Iturrieta-González, Dania García & Gené; Alternaria pobletensis Iturrieta-González, Dania García & Gené; Alternaria pseudoventricosa Iturrieta-González, Dania García & Gené; Arezzomyces Y. Marín & Crous; Arezzomyces cytisi (Wanas. et al.) Y. Marín & Crous; Ascochyta chrysanthemi F. Stevens; Brunneosphaerella roupeliae Crous; DNA barcodes; Elsinoe picconiae Crous; Elsinoe veronicae Crous, Thangavel & Y. Marín; Fungal systematics; Globoramichloridium Y. Marín & Crous; Globoramichloridium indicum (Subram.) Y. Marín & Crous; Neosetophoma aseptata Crous, R.K. Schumach. & Y. Marín; Neosetophoma phragmitis Crous, R.K. Schumach. & Y. Marín; Neosetophoma sambuci Crous, R.K. Schumach. & Y. Marín; Neostagonospora sorghi Crous & Y. Marín; New taxa; Parastagonospora novozelandica Crous, Thangavel & Y. Marín; Parastagonospora phragmitis Crous & Y. Marín; Pestalotia unicornis Cooke & Ellis; Phaeosphaeria phoenicicola (Crous & Thangavel) Y. Marín & Crous; Phaeosphaeriopsis aloes Crous & Y. Marín; Phaeosphaeriopsis aloicola Crous & Y. Marín; Phaeosphaeriopsis grevilleae Crous & Y. Marín; Phaeosphaeriopsis pseudoagavacearum Crous & Y. Marín; Pleiocarpon livistonae Crous & Quaedvl.; Pyrenophora avenicola Y. Marín & Crous; Pyrenophora cynosuri Y. Marín & Crous; Pyrenophora nisikadoi Y. Marín & Crous; Pyrenophora novozelandica Y. Marín & Crous; Pyrenophora poae (Baudyš) Y. Marín & Crous; Pyrenophora pseudoerythrospila Y. Marín & Crous; Pyrenophora sieglingiae Y. Marín & Crous; Pyrenophora variabilis Hern.-Restr. & Y. Marín; Pyrenophora wirreganensis (Wallwork et al.) Y. Marín & Crous; Rhynchosphaeria cupressi Nattrass et al; Seiridium cupressi (Nattrass et al.) Bonthond, Sandoval-Denis & Crous; Seiridium pezizoides (de Not.) Crous; Septoriella agrostina (Mapook et al.) Y. Marín & Crous; Septoriella artemisiae (Wanas. et al.) Y. Marín & Crous; Septoriella arundinicola (Wanas. et al.) Y. Marín & Crous; Septoriella arundinis (W.J. Li et al.) Y. Marín & Crous; Septoriella bromi (Wijayaw. et al.) Y. Marín & Crous; Septoriella dactylidicola Y. Marín & Crous; Septoriella dactylidis (Wanas. et al.) Y. Marín & Crous; Septoriella elongata (Wehm.) Y. Marín & Crous; Septoriella forlicesenica (Thambug. et al.) Y. Marín & Crous; Septoriella garethjonesii (Thambug. et al.) Y. Marín & Crous; Septoriella germanica Crous, R.K. Schumach. & Y. Marín; Septoriella hibernica Crous, Quaedvl. & Y. Marín; Septoriella hollandica Crous, Quaedvl. & Y. Marín; Septoriella italica (Thambug. et al.) Y. Marín & Crous; Septoriella muriformis (Ariyaw. et al.) Y. Marín & Crous; Septoriella neoarundinis Y. Marín & Crous; Septoriella neodactylidis Y. Marín & Crous; Septoriella pseudophragmitis Crous, Quaedvl. & Y. Marín; Septoriella rosae (Mapook et al.) Y. Marín & Crous; Septoriella subcylindrospora (W.J. Li et al.) Y. Marín & Crous; Septoriella vagans (Niessl) Y. Marín & Crous; Setophoma brachypodii Crous, R.K. Schumach. & Y. Marín; Setophoma pseudosacchari Crous & Y. Marín; Stemphylium rombundicum Moslemi, Y.P. Tan & P.W.J. Taylor; Stemphylium truncatulae Moslemi, Y.P. Tan & P.W.J. Taylor; Stemphylium waikerieanum Moslemi, Jacq. Edwards & P.W.J Taylor; Vagicola arundinis Phukhams., Camporesi & K.D. Hyde; Wingfieldomyces Y. Marín & Crous; Wingfieldomyces cyperi (Crous & M.J. Wingf.) Y. Marín & Crous; Zasmidium ducassei (R.G. Shivas et al.) Y. Marín & Crous; Zasmidium thailandicum Crous.
© 2019 Westerdijk Fungal Biodiversity Institute. Production and hosting by ELSEVIER B.V.
Figures







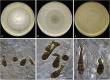



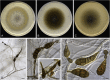


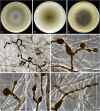























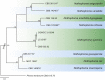





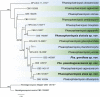























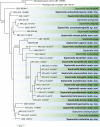













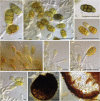





References
-
- Adler A., Yaniv I., Samra Z. Exserohilum: an emerging human pathogen. European Journal of Clinical Microbiology and Infectious Diseases. 2006;25:247–253. - PubMed
-
- Ahmed S.A., Hofmüller W., Seibold M. Tintelnotia, a new genus in Phaeosphaeriaceae harbouring agents of cornea and nail infections in humans. Mycoses. 2017;60:244–253. - PubMed
-
- Amaradasa B.S., Madrid H., Groenewald J.Z. Porocercospora seminalis gen. et comb. nov. the causal organism of buffalo grass false smut. Mycologia. 2014;106:77–85. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
