Transcriptome analysis offers a comprehensive illustration of the genetic background of pediatric acute myeloid leukemia
- PMID: 31648321
- PMCID: PMC6849955
- DOI: 10.1182/bloodadvances.2019000404
Transcriptome analysis offers a comprehensive illustration of the genetic background of pediatric acute myeloid leukemia
Abstract
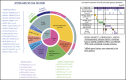
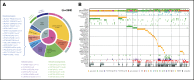
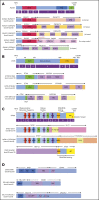
Recent advances in the genetic understanding of acute myeloid leukemia (AML) have improved clinical outcomes in pediatric patients. However, ∼40% of patients with pediatric AML relapse, resulting in a relatively low overall survival rate of ∼70%. The objective of this study was to reveal the comprehensive genetic background of pediatric AML. We performed transcriptome analysis (RNA sequencing [RNA-seq]) in 139 of the 369 patients with de novo pediatric AML who were enrolled in the Japanese Pediatric Leukemia/Lymphoma Study Group AML-05 trial and investigated correlations between genetic aberrations and clinical information. Using RNA-seq, we identified 54 in-frame gene fusions and 1 RUNX1 out-of-frame fusion in 53 of 139 patients. Moreover, we found at least 258 gene fusions in 369 patients (70%) through reverse transcription polymerase chain reaction and RNA-seq. Five gene rearrangements were newly identified, namely, NPM1-CCDC28A, TRIP12-NPM1, MLLT10-DNAJC1, TBL1XR1-RARB, and RUNX1-FNBP1. In addition, we found rare gene rearrangements, namely, MYB-GATA1, NPM1-MLF1, ETV6-NCOA2, ETV6-MECOM, ETV6-CTNNB1, RUNX1-PRDM16, RUNX1-CBFA2T2, and RUNX1-CBFA2T3. Among the remaining 111 patients, KMT2A-PTD, biallelic CEBPA, and NPM1 gene mutations were found in 11, 23, and 17 patients, respectively. These mutations were completely mutually exclusive with any gene fusions. RNA-seq unmasked the complexity of gene rearrangements and mutations in pediatric AML. We identified potentially disease-causing alterations in nearly all patients with AML, including novel gene fusions. Our results indicated that a subset of patients with pediatric AML represent a distinct entity that may be discriminated from their adult counterparts. Based on these results, risk stratification should be reconsidered.
© 2019 by The American Society of Hematology.
Conflict of interest statement
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Figures



References
-
- Fröhling S, Scholl C, Gilliland DG, Levine RL. Genetics of myeloid malignancies: pathogenetic and clinical implications. J Clin Oncol. 2005;23(26):6285-6295. - PubMed
-
- Marcucci G, Haferlach T, Döhner H. Molecular genetics of adult acute myeloid leukemia: prognostic and therapeutic implications. J Clin Oncol. 2011;29(5):475-486. - PubMed
-
- Preudhomme C, Sagot C, Boissel N, et al. ; ALFA Group. Favorable prognostic significance of CEBPA mutations in patients with de novo acute myeloid leukemia: a study from the Acute Leukemia French Association (ALFA). Blood. 2002;100(8):2717-2723. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

