miR-29a Is Repressed by MYC in Pancreatic Cancer and Its Restoration Drives Tumor-Suppressive Effects via Downregulation of LOXL2
- PMID: 31662451
- PMCID: PMC7002266
- DOI: 10.1158/1541-7786.MCR-19-0594
miR-29a Is Repressed by MYC in Pancreatic Cancer and Its Restoration Drives Tumor-Suppressive Effects via Downregulation of LOXL2
Abstract
Pancreatic ductal adenocarcinoma (PDAC) is an intractable cancer with a dismal prognosis. miR-29a is commonly downregulated in PDAC; however, mechanisms for its loss and role still remain unclear. Here, we show that in PDAC, repression of miR-29a is directly mediated by MYC via promoter activity. RNA sequencing analysis, integrated with miRNA target prediction, identified global miR-29a downstream targets in PDAC. Target enrichment coupled with gene ontology and survival correlation analyses identified the top five miR-29a-downregulated target genes (LOXL2, MYBL2, CLDN1, HGK, and NRAS) that are known to promote tumorigenic mechanisms. Functional validation confirmed that upregulation of miR-29a is sufficient to ablate translational expression of these five genes in PDAC. We show that the most promising target among the identified genes, LOXL2, is repressed by miR-29a via 3'-untranslated region binding. Pancreatic tissues from a PDAC murine model and patient biopsies showed overall high LOXL2 expression with inverse correlations with miR-29a levels. Collectively, our data delineate an antitumorigenic, regulatory role of miR-29a and a novel MYC-miR-29a-LOXL2 regulatory axis in PDAC pathogenesis, indicating the potential of the molecule in therapeutic opportunities. IMPLICATIONS: This study unravels a novel functional role of miR-29a in PDAC pathogenesis and identifies an MYC-miR-29a-LOXL2 axis in regulation of the disease progression, implicating miR-29a as a potential therapeutic target for PDAC. VISUAL OVERVIEW: http://mcr.aacrjournals.org/content/molcanres/18/2/311/F1.large.jpg.
©2019 American Association for Cancer Research.
Conflict of interest statement
Figures



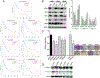
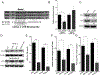

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

