METTL15 introduces N4-methylcytidine into human mitochondrial 12S rRNA and is required for mitoribosome biogenesis
- PMID: 31665743
- PMCID: PMC6821322
- DOI: 10.1093/nar/gkz735
METTL15 introduces N4-methylcytidine into human mitochondrial 12S rRNA and is required for mitoribosome biogenesis
Abstract
Post-transcriptional RNA modifications, the epitranscriptome, play important roles in modulating the functions of RNA species. Modifications of rRNA are key for ribosome production and function. Identification and characterization of enzymes involved in epitranscriptome shaping is instrumental for the elucidation of the functional roles of specific RNA modifications. Ten modified sites have been thus far identified in the mammalian mitochondrial rRNA. Enzymes responsible for two of these modifications have not been characterized. Here, we identify METTL15, show that it is the main N4-methylcytidine (m4C) methyltransferase in human cells and demonstrate that it is responsible for the methylation of position C839 in mitochondrial 12S rRNA. We show that the lack of METTL15 results in a reduction of the mitochondrial de novo protein synthesis and decreased steady-state levels of protein components of the oxidative phosphorylation system. Without functional METTL15, the assembly of the mitochondrial ribosome is decreased, with the late assembly components being unable to be incorporated efficiently into the small subunit. We speculate that m4C839 is involved in the stabilization of 12S rRNA folding, therefore facilitating the assembly of the mitochondrial small ribosomal subunits. Taken together our data show that METTL15 is a novel protein necessary for efficient translation in human mitochondria.
© The Author(s) 2019. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

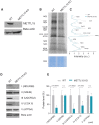





References
-
- Greber B.J., Boehringer D., Leibundgut M., Bieri P., Leitner A., Schmitz N., Aebersold R., Ban N.. The complete structure of the large subunit of the mammalian mitochondrial ribosome. Nature. 2014; 515:283–286. - PubMed
-
- Greber B.J., Boehringer D., Leitner A., Bieri P., Voigts-Hoffmann F., Erzberger J.P., Leibundgut M., Aebersold R., Ban N.. Architecture of the large subunit of the mammalian mitochondrial ribosome. Nature. 2014; 505:515–519. - PubMed
-
- Greber B.J., Bieri P., Leibundgut M., Leitner A., Aebersold R., Boehringer D., Ban N.. Ribosome. The complete structure of the 55S mammalian mitochondrial ribosome. Science. 2015; 348:303–308. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

