Genome replication dynamics of a bacteriophage and its satellite reveal strategies for parasitism and viral restriction
- PMID: 31667508
- PMCID: PMC7145576
- DOI: 10.1093/nar/gkz1005
Genome replication dynamics of a bacteriophage and its satellite reveal strategies for parasitism and viral restriction
Abstract
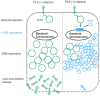
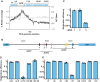
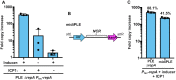
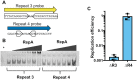
Phage-inducible chromosomal island-like elements (PLEs) are bacteriophage satellites found in Vibrio cholerae. PLEs parasitize the lytic phage ICP1, excising from the bacterial chromosome, replicating, and mobilizing to new host cells following cell lysis. PLEs protect their host cell populations by completely restricting the production of ICP1 progeny. Previously, it was found that ICP1 replication was reduced during PLE(+) infection. Despite robust replication of the PLE genome, relatively few transducing units are produced. We investigated if PLE DNA replication itself is antagonistic to ICP1 replication. Here we identify key constituents of PLE replication and assess their role in interference of ICP1. PLE encodes a RepA_N initiation factor that is sufficient to drive replication from the PLE origin of replication during ICP1 infection. In contrast to previously characterized bacteriophage satellites, expression of the PLE initiation factor was not sufficient for PLE replication in the absence of phage. Replication of PLE was necessary for interference of ICP1 DNA replication, but replication of a minimalized PLE replicon was not sufficient for ICP1 DNA replication interference. Despite restoration of ICP1 DNA replication, non-replicating PLE remained broadly inhibitory against ICP1. These results suggest that PLE DNA replication is one of multiple mechanisms contributing to ICP1 restriction.
© The Author(s) 2019. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


Similar articles
-
A Family of Viral Satellites Manipulates Invading Virus Gene Expression and Can Affect Cholera Toxin Mobilization.mSystems. 2020 Oct 13;5(5):e00358-20. doi: 10.1128/mSystems.00358-20. mSystems. 2020. PMID: 33051375 Free PMC article.
-
Viral Satellites Exploit Phage Proteins to Escape Degradation of the Bacterial Host Chromosome.Cell Host Microbe. 2019 Oct 9;26(4):504-514.e4. doi: 10.1016/j.chom.2019.09.006. Cell Host Microbe. 2019. PMID: 31600502 Free PMC article.
-
A chimeric nuclease substitutes a phage CRISPR-Cas system to provide sequence-specific immunity against subviral parasites.Elife. 2021 Jul 7;10:e68339. doi: 10.7554/eLife.68339. Elife. 2021. PMID: 34232860 Free PMC article.
-
Multiple regulatory mechanisms controlling phage-plasmid P4 propagation.FEMS Microbiol Rev. 1995 Aug;17(1-2):127-34. doi: 10.1111/j.1574-6976.1995.tb00194.x. FEMS Microbiol Rev. 1995. PMID: 7669338 Review.
-
Coevolution of bacteria and their viruses.Folia Microbiol (Praha). 2013 May;58(3):177-86. doi: 10.1007/s12223-012-0195-5. Epub 2012 Sep 21. Folia Microbiol (Praha). 2013. PMID: 22993102 Review.
Cited by
-
Phage-inducible chromosomal minimalist islands (PICMIs), a novel family of small marine satellites of virulent phages.Nat Commun. 2024 Jan 22;15(1):664. doi: 10.1038/s41467-024-44965-1. Nat Commun. 2024. PMID: 38253718 Free PMC article.
-
Bacteriophage ICP1: A Persistent Predator of Vibrio cholerae.Annu Rev Virol. 2021 Sep 29;8(1):285-304. doi: 10.1146/annurev-virology-091919-072020. Epub 2021 Jul 27. Annu Rev Virol. 2021. PMID: 34314595 Free PMC article.
-
A Family of Viral Satellites Manipulates Invading Virus Gene Expression and Can Affect Cholera Toxin Mobilization.mSystems. 2020 Oct 13;5(5):e00358-20. doi: 10.1128/mSystems.00358-20. mSystems. 2020. PMID: 33051375 Free PMC article.
-
Fine-tuned spatiotemporal dynamics of DNA replication during phage lambda infection.J Virol. 2024 Nov 19;98(11):e0112824. doi: 10.1128/jvi.01128-24. Epub 2024 Oct 31. J Virol. 2024. PMID: 39480083 Free PMC article.
-
Nuclease genes occupy boundaries of genetic exchange between bacteriophages.NAR Genom Bioinform. 2023 Aug 24;5(3):lqad076. doi: 10.1093/nargab/lqad076. eCollection 2023 Sep. NAR Genom Bioinform. 2023. PMID: 37636022 Free PMC article.
References
-
- Krupovic M., Kuhn J.H., Fischer M.G.. A classification system for virophages and satellite viruses. Arch. Virol. 2016; 161:233–247. - PubMed
-
- Contursi P., Fusco S., Cannio R., She Q.. Molecular biology of fuselloviruses and their satellites. Extremophiles. 2014; 18:473–489. - PubMed
-
- Hughes S.A., Wedemeyer H., Harrison P.M.. Hepatitis delta virus. Lancet. 2011; 378:73–85. - PubMed
-
- Simon A.E., Roossinck M.J., Havelda Z.. Plant virus satellite and defective interfering RNAs: new paradigms for a new century. Annu. Rev. Phytopathol. 2004; 42:415–437. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

