Intra-Patient Heterogeneity of Circulating Tumor Cells and Circulating Tumor DNA in Blood of Melanoma Patients
- PMID: 31671846
- PMCID: PMC6896052
- DOI: 10.3390/cancers11111685
Intra-Patient Heterogeneity of Circulating Tumor Cells and Circulating Tumor DNA in Blood of Melanoma Patients
Abstract
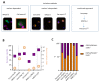
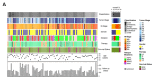
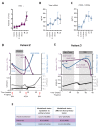
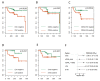
Despite remarkable progress in melanoma therapy, the exceptional heterogeneity of the disease has prevented the development of reliable companion biomarkers for the prediction or monitoring of therapy responses. Here, we show that difficulties in detecting blood-based markers, like circulating tumor cells (CTC), might arise from the translation of the mutational heterogeneity of melanoma cells towards their surface marker expression. We provide a unique method, which enables the molecular characterization of clinically relevant CTC subsets, as well as circulating tumor DNA (ctDNA), from a single blood sample. The study demonstrates the benefit of a combined analysis of ctDNA and CTC counts in melanoma patients, revealing that CTC subsets and ctDNA provide synergistic real-time information on the mutational status, RNA and protein expression of melanoma cells in individual patients, in relation to clinical outcome.
Keywords: CTC; ctDNA; liquid biopsy; melanoma.
Conflict of interest statement
A. Sartori is employed by Agena Bioscience GmbH. The authors declare no conflict of interest.
Figures


References
Grants and funding
LinkOut - more resources
Full Text Sources

