Twist1-Induced Epithelial Dissemination Requires Prkd1 Signaling
- PMID: 31676574
- PMCID: PMC7448227
- DOI: 10.1158/0008-5472.CAN-18-3241
Twist1-Induced Epithelial Dissemination Requires Prkd1 Signaling
Abstract
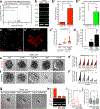
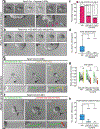
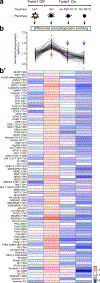
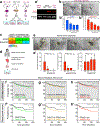
Dissemination is an essential early step in metastasis but its molecular basis remains incompletely understood. To define the essential targetable effectors of this process, we developed a 3D mammary epithelial culture model, in which dissemination is induced by overexpression of the transcription factor Twist1. Transcriptomic analysis and ChIP-PCR together demonstrated that protein kinase D1 (Prkd1) is a direct transcriptional target of Twist1 and is not expressed in the normal mammary epithelium. Pharmacologic and genetic inhibition of Prkd1 in the Twist1-induced dissemination model demonstrated that Prkd1 was required for cells to initiate extracellular matrix (ECM)-directed protrusions, release from the epithelium, and migrate through the ECM. Antibody-based protein profiling revealed that Prkd1 induced broad phosphorylation changes, including an inactivating phosphorylation of β-catenin and two microtubule depolymerizing phosphorylations of Tau, potentially explaining the release of cell-cell contacts and persistent activation of Prkd1. In patients with breast cancer, TWIST1 and PRKD1 expression correlated with metastatic recurrence, particularly in basal breast cancer. Prkd1 knockdown was sufficient to block dissemination of both murine and human mammary tumor organoids. Finally, Prkd1 knockdown in vivo blocked primary tumor invasion and distant metastasis in a mouse model of basal breast cancer. Collectively, these data identify Prkd1 as a novel and targetable signaling node downstream of Twist1 that is required for epithelial invasion and dissemination. SIGNIFICANCE: Twist1 is a known regulator of metastatic cell behaviors but not directly targetable. This study provides a molecular explanation for how Twist1-induced dissemination works and demonstrates that it can be targeted. GRAPHICAL ABSTRACT: http://cancerres.aacrjournals.org/content/canres/80/2/204/F1.large.jpg.
©2019 American Association for Cancer Research.
Conflict of interest statement
Figures


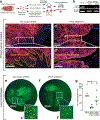
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

