Regulation of Pyridine Nucleotide Metabolism During Tomato Fruit Development Through Transcript and Protein Profiling
- PMID: 31681351
- PMCID: PMC6798084
- DOI: 10.3389/fpls.2019.01201
Regulation of Pyridine Nucleotide Metabolism During Tomato Fruit Development Through Transcript and Protein Profiling
Abstract
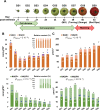
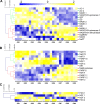
Central metabolism is the engine of plant biomass, supplying fruit growth with building blocks, energy, and biochemical cofactors. Among metabolic cornerstones, nicotinamide adenine dinucleotide (NAD) is particularly pivotal for electron transfer through reduction-oxidation (redox) reactions, thus participating in a myriad of biochemical processes. Besides redox functions, NAD is now assumed to act as an integral regulator of signaling cascades involved in growth and environmental responses. However, the regulation of NAD metabolism and signaling during fruit development remains poorly studied and understood. Here, we benefit from RNAseq and proteomic data obtained from nine growth stages of tomato fruit (var. Moneymaker) to dissect mRNA and protein profiles that link to NAD metabolism, including de novo biosynthesis, recycling, utilization, and putative transport. As expected for a cofactor synthesis pathway, protein profiles failed to detect enzymes involved in NAD synthesis or utilization, except for nicotinic acid phosphoribosyltransferase (NaPT) and nicotinamidase (NIC), which suggested that most NAD metabolic enzymes were poorly represented quantitatively. Further investigations on transcript data unveiled differential expression patterns during fruit development. Interestingly, among specific NAD metabolism-related genes, early de novo biosynthetic genes were transcriptionally induced in very young fruits, in association with NAD kinase, while later stages of fruit growth rather showed an accumulation of transcripts involved in later stages of de novo synthesis and in NAD recycling, which agreed with augmented NAD(P) levels. In addition, a more global overview of 119 mRNA and 78 protein significant markers for NAD(P)-dependent enzymes revealed differential patterns during tomato growth that evidenced clear regulations of primary metabolism, notably with respect to mitochondrial functions. Overall, we propose that NAD metabolism and signaling are very dynamic in the developing tomato fruit and that its differential regulation is certainly critical to fuel central metabolism linking to growth mechanisms.
Keywords: NAD; development; fruit; redox; tomato.
Copyright © 2019 Decros, Beauvoit, Colombié, Cabasson, Bernillon, Arrivault, Guenther, Belouah, Prigent, Baldet, Gibon and Pétriacq.
Figures



References
-
- Araújo W. L., Nunes-Nesi A., Nikoloski Z., Sweetlove L. J., Fernie A. R. (2012). Metabolic control and regulation of the tricarboxylic acid cycle in photosynthetic and heterotrophic plant tissues: TCA control and regulation in plant tissues. Plant Cell Environ. 35, 1–21. 10.1111/j.1365-3040.2011.02332.x - DOI - PubMed
-
- Arrivault S., Guenther M., Ivakov A., Feil R., Vosloh D., van Dongen J. T., et al. (2009). Use of reverse-phase liquid chromatography, linked to tandem mass spectrometry, to profile the Calvin cycle and other metabolic intermediates in Arabidopsis rosettes at different carbon dioxide concentrations. Plant J. 59, 826–839. 10.1111/j.1365-313X.2009.03902.x - DOI - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

