Discovery of holoenzyme-disrupting chemicals as substrate-selective CK2 inhibitors
- PMID: 31685885
- PMCID: PMC6828666
- DOI: 10.1038/s41598-019-52141-5
Discovery of holoenzyme-disrupting chemicals as substrate-selective CK2 inhibitors
Abstract
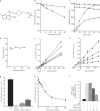
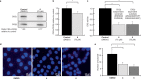
CK2 is a constitutively active protein kinase overexpressed in numerous malignancies. Interaction between CK2α and CK2β subunits is essential for substrate selectivity. The CK2α/CK2β interface has been previously targeted by peptides to achieve functional effects; however, no small molecules modulators were identified due to pocket flexibility and open shape. Here we generated numerous plausible conformations of the interface using the fumigation modeling protocol, and virtually screened a compound library to discover compound 1 that suppressed CK2α/CK2β interaction in vitro and inhibited CK2 in a substrate-selective manner. Orthogonal SPR, crystallography, and NMR experiments demonstrated that 4 and 6, improved analogs of 1, bind to CK2α as predicted. Both inhibitors alter CK2 activity in cells through inhibition of CK2 holoenzyme formation. Treatment with 6 suppressed MDA-MB231 triple negative breast cancer cell growth and induced apoptosis. Altogether, our findings exemplify an innovative computational-experimental approach and identify novel non-peptidic inhibitors of CK2 subunit interface disclosing substrate-selective functional effects.
Conflict of interest statement
R.A. has an equity interest in Molsoft, LLC. The terms of this arrangement have been reviewed and approved by the University of California, San Diego in accordance with its conflict of interest policies.
Figures