High-resolution ion mobility spectrometry-mass spectrometry of isomeric/isobaric ribonucleotide variants
- PMID: 31697854
- PMCID: PMC8363168
- DOI: 10.1002/jms.4465
High-resolution ion mobility spectrometry-mass spectrometry of isomeric/isobaric ribonucleotide variants
Abstract
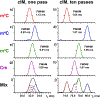
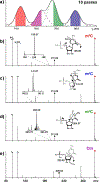
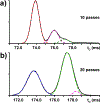
In this report, we explored the benefits of cyclic ion mobility (cIM) mass spectrometry in the analysis of isomeric post-transcriptional modifications of RNA. Standard methyl-cytidine samples were initially utilized to test the ability to correctly distinguish different structures sharing the same elemental composition and thus molecular mass. Analyzed individually, the analytes displayed characteristic arrival times (tD ) determined by the different positions of the modifying methyl groups onto the common cytidine scaffold. Analyzed in mixture, the widths of the respective signals resulted in significant overlap that initially prevented their resolution on the tD scale. The separation of the four isomers was achieved by increasing the number of passes through the cIM device, which enabled to fully differentiate the characteristic ion mobility behaviors associated with very subtle structural variations. The placement of the cIM device between the mass-selective quadrupole and the time-of-flight analyzer allowed us to perform gas-phase activation of each of these ion populations, which had been first isolated according to a common mass-to-charge ratio and then separated on the basis of different ion mobility behaviors. The observed fragmentation patterns confirmed the structures of the various isomers thus substantiating the benefits of complementing unique tD information with specific fragmentation data to reach more stringent analyte identification. These capabilities were further tested by analyzing natural mono-nucleotide mixtures obtained by exonuclease digestion of total RNA extracts. In particular, the combination of cIM separation and post-mobility dissociation allowed us to establish the composition of methyl-cytidine and methyl-adenine components present in the entire transcriptome of HeLa cells. For this reason, we expect that this technique will benefit not only epitranscriptomic studies requiring the determination of identity and expression levels of RNA modifications, but also metabolomics investigations involving the analysis of natural extracts that may possibly contain subsets of isomeric/isobaric species.
© 2019 John Wiley & Sons, Ltd.
Conflict of interest statement
Conflict of interest statement. None declared.
Figures
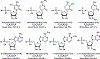




References
-
- Grosjean H: Fine-tuning of RNA functions by modification and editing. (Springer, 2005).
-
- Carell T, Brandmayr C, Hienzsch A, Müller M, Pearson D, Reiter V, Thoma I, Thumbs P. & Wagner M: Structure and function of noncanonical nucleobases. Angew. Chem. Int. Ed. Engl 51, 7110–7131 (2012). - PubMed
-
- Mattick JS & Makunin IV: Non-coding RNA. Human molecular genetics 15 Spec No 1, R17–29 (2006). - PubMed
-
- Shafik A, Schumann U, Evers M, Sibbritt T. & Preiss T: The emerging epitranscriptomics of long noncoding RNAs. Biochim. Biophys. Acta 1859, 59–70 (2015). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

