HIV‑1 integrase inhibitors targeting various DDE transposases: Retroviral integration versus RAG‑mediated recombination (Review)
- PMID: 31702817
- PMCID: PMC6854553
- DOI: 10.3892/mmr.2019.10777
HIV‑1 integrase inhibitors targeting various DDE transposases: Retroviral integration versus RAG‑mediated recombination (Review)
Abstract
Transposases are ubiquitous mobile genetic elements responsible for genome development, driving rearrangements, such as insertions, deletions and translocations. Across species evolution, some transposases are tamed by their host and are made part of complex cellular systems. The proliferation of retroviruses is also dependent on transposase related enzymes termed integrases. Recombination‑activating gene protein (RAG)1 and metnase are just two examples of transposase domestication and together with retroviral integrases (INs), they belong to the DDE polynucleotidyl transferases superfamily. They share mechanistic and structural features linked to the RNase H‑like fold, harboring a DDE(D) metal dependent catalytic motif. Recent antiretroviral compounds target the catalytic domain of integrase, but they also have the potential of inhibiting other related enzymes. In this review, we report the activity of different classes of integrase inhibitors on various DDE transposases. Computational simulations are useful to predict the extent of off‑target activity and have been employed to study the interactions between RAG1 recombinase and compounds from three different pharmacologic classes. We demonstrate that strand‑transfer inhibitors display a higher affinity towards the RAG1 RNase H domain, as suggested by experimental data compared to allosteric inhibitors. While interference with RAG1 and 2 recombination is associated with a negative impact on immune function, the inhibition of metnase or HTLV‑1 integrase opens the way for the development of novel therapies for refractory cancers.
Figures



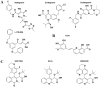



References
-
- Lee SH, Oshige M, Durant ST, Rasila KK, Williamson EA, Ramsey H, Kwan L, Nickoloff JA, Hromas R. The SET domain protein Metnase mediates foreign DNA integration and links integration to nonhomologous end-joining repair. Proc Natl Acad Sci USA. 2005;102:18075–18080. doi: 10.1073/pnas.0503676102. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

