Mouse APOBEC1 cytidine deaminase can induce somatic mutations in chromosomal DNA
- PMID: 31726973
- PMCID: PMC6854741
- DOI: 10.1186/s12864-019-6216-x
Mouse APOBEC1 cytidine deaminase can induce somatic mutations in chromosomal DNA
Abstract
Background: APOBEC1 (A1) enzymes are cytidine deaminases involved in RNA editing. In addition to this activity, a few A1 enzymes have been shown to be active on single stranded DNA. As two human ssDNA cytidine deaminases APOBEC3A (A3A), APOBEC3B (A3B) and related enzymes across the spectrum of placental mammals have been shown to introduce somatic mutations into nuclear DNA of cancer genomes, we explored the mutagenic threat of A1 cytidine deaminases to chromosomal DNA.
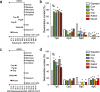
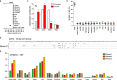
Results: Molecular cloning and expression of various A1 enzymes reveal that the cow, pig, dog, rabbit and mouse A1 have an intracellular ssDNA substrate specificity. However, among all the enzymes studied, mouse A1 appears to be singular, being able to introduce somatic mutations into nuclear DNA with a clear 5'TpC editing context, and to deaminate 5-methylcytidine substituted DNA which are characteristic features of the cancer related mammalian A3A and A3B enzymes. However, mouse A1 activity fails to elicit formation of double stranded DNA breaks, suggesting that mouse A1 possess an attenuated nuclear DNA mutator phenotype reminiscent of human A3B.
Conclusions: At an experimental level mouse APOBEC1 is remarkable among 12 mammalian A1 enzymes in that it represents a source of somatic mutations in mouse genome, potentially fueling oncogenesis. While the order Rodentia is bereft of A3A and A3B like enzymes it seems that APOBEC1 may well substitute for it, albeit remaining much less active. This modifies the paradigm that APOBEC3 and AID enzymes are the sole endogenous mutator enzymes giving rise to off-target editing of mammalian genomes.
Keywords: APOBEC1; Cancer; Cytidine deaminase; Nuclear DNA; Somatic mutations.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



References
-
- Navaratnam N, Morrison JR, Bhattacharya S, Patel D, Funahashi T, Giannoni F, et al. The p27 catalytic subunit of the apolipoprotein B mRNA editing enzyme is a cytidine deaminase. J Biol Chem. 1993;268:20709–20712. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

