Improved risk-stratification for posterior fossa ependymoma of childhood considering clinical, histological and genetic features - a retrospective analysis of the HIT ependymoma trial cohort
- PMID: 31727173
- PMCID: PMC6857225
- DOI: 10.1186/s40478-019-0820-5
Improved risk-stratification for posterior fossa ependymoma of childhood considering clinical, histological and genetic features - a retrospective analysis of the HIT ependymoma trial cohort
Abstract
Introduction: Risk stratification of children with ependymomas of the posterior fossa in current therapeutic protocols is mainly based on clinical criteria. We aimed to identify independent outcome predictors for this disease entity by a systematic integrated analysis of clinical, histological and genetic information in a defined cohort of patients treated according to the German HIT protocols.
Methods: Tumor samples of 134 patients aged 0.2-15.9 years treated between 1999 and 2010 according to HIT protocols were analyzed for histological features including mitotic activity, necrosis and vascular proliferation and genomic alterations by SNP and molecular inversion probe analysis. Survival analysis was performed by Kaplan-Meier method with log rank test and multivariate Cox regression analysis.
Results: Residual tumor after surgery, chromosome 1q gain and structural genomic alterations were identified as predictors of significantly shorter event-free (EFS) and overall survival (OS). Furthermore, specific histological features including vascular proliferation, necrosis and high mitotic activity were predictive for shorter OS. Multivariate Cox regression revealed residual tumor, chromosome 1q gain and mitotic activity as independent predictors of both EFS and OS. Using these independent predictors of outcome, we were able to build a 3-tiered risk stratification model that separates patients with standard, intermediate and high risk, and which outperforms current stratification procedures.
Conclusion: The integration of defined clinical, histological and genetic parameters led to an improved risk-stratification model for posterior fossa ependymoma of childhood. After validation in independent cohorts this model may provide the basis for risk-adapted treatment of children with ependymomas of the posterior fossa.
Keywords: Ependymoma; Genetics; Neuropathology; Posterior fossa; Risk stratification.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
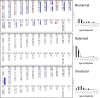


References
-
- Andreiuolo F, Varlet P, Tauziede-Espariat A, Junger ST, Dorner E, Dreschmann V, Kuchelmeister K, Waha A, Haberler C, Slavc I, Corbacioglu S, Riemenschneider MJ, Leipold A, Rudiger T, Korholz D, Acker T, Russo A, Faber J, Sommer C, Armbrust S, Rose M, Erdlenbruch B, Hans VH, Bernbeck B, Schneider D, Lorenzen J, Ebinger M, Handgretinger R, Neumann M, van Buiren M, Prinz M, Roganovic J, Jakovcevic A, Park SH, Grill J, Puget S, Messing-Junger M, Reinhard H, Bergmann M, Hattingen E, Pietsch T (2018) Childhood supratentorial ependymomas with YAP1-MAMLD1 fusion: an entity with characteristic clinical, radiological, cytogenetic and histopathological features. Brain Pathol. 10.1111/bpa.12659 - PubMed
-
- Beroukhim R, Getz G, Nghiemphu L, Barretina J, Hsueh T, Linhart D, Vivanco I, Lee JC, Huang JH, Alexander S, Du J, Kau T, Thomas RK, Shah K, Soto H, Perner S, Prensner J, Debiasi RM, Demichelis F, Hatton C, Rubin MA, Garraway LA, Nelson SF, Liau L, Mischel PS, Cloughesy TF, Meyerson M, Golub TA, Lander ES, Mellinghoff IK, Sellers WR. Assessing the significance of chromosomal aberrations in cancer: methodology and application to glioma. Proc Natl Acad Sci U S A. 2007;104:20007–20012. doi: 10.1073/pnas.0710052104. - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

