Disease Associated Mutations in KIR Proteins Linked to Aberrant Inward Rectifier Channel Trafficking
- PMID: 31731488
- PMCID: PMC6920955
- DOI: 10.3390/biom9110650
Disease Associated Mutations in KIR Proteins Linked to Aberrant Inward Rectifier Channel Trafficking
Abstract
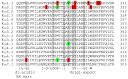
The ubiquitously expressed family of inward rectifier potassium (KIR) channels, encoded by KCNJ genes, is primarily involved in cell excitability and potassium homeostasis. Channel mutations associate with a variety of severe human diseases and syndromes, affecting many organ systems including the central and peripheral neural system, heart, kidney, pancreas, and skeletal muscle. A number of mutations associate with altered ion channel expression at the plasma membrane, which might result from defective channel trafficking. Trafficking involves cellular processes that transport ion channels to and from their place of function. By alignment of all KIR channels, and depicting the trafficking associated mutations, three mutational hotspots were identified. One localized in the transmembrane-domain 1 and immediately adjacent sequences, one was found in the G-loop and Golgi-export domain, and the third one was detected at the immunoglobulin-like domain. Surprisingly, only few mutations were observed in experimentally determined Endoplasmic Reticulum (ER)exit-, export-, or ER-retention motifs. Structural mapping of the trafficking defect causing mutations provided a 3D framework, which indicates that trafficking deficient mutations form clusters. These "mutation clusters" affect trafficking by different mechanisms, including protein stability.
Keywords: KCNJ; KIR; alignment; disease; inward rectifier channel; mutation; structure; trafficking.
Conflict of interest statement
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures




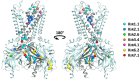

References
-
- Katz B. Les constantes électriques de la membrane du muscle. Arch. Sci. Physiol. 1949;2:285–299.
-
- Wang L., Chiamvimonvat N., Duff H.J. Interaction between selected sodium and potassium channel blockers in guinea pig papillary muscle. J. Pharmacol. Exp. Ther. 1993;264:1056–1062. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

