Immunopeptidomics of colorectal cancer organoids reveals a sparse HLA class I neoantigen landscape and no increase in neoantigens with interferon or MEK-inhibitor treatment
- PMID: 31735170
- PMCID: PMC6859637
- DOI: 10.1186/s40425-019-0769-8
Immunopeptidomics of colorectal cancer organoids reveals a sparse HLA class I neoantigen landscape and no increase in neoantigens with interferon or MEK-inhibitor treatment
Abstract
Background: Patient derived organoids (PDOs) can be established from colorectal cancers (CRCs) as in vitro models to interrogate cancer biology and its clinical relevance. We applied mass spectrometry (MS) immunopeptidomics to investigate neoantigen presentation and whether this can be augmented through interferon gamma (IFNγ) or MEK-inhibitor treatment.
Methods: Four microsatellite stable PDOs from chemotherapy refractory and one from a treatment naïve CRC were expanded to replicates with 100 million cells each, and HLA class I and class II peptide ligands were analyzed by MS.
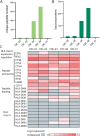
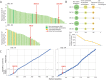
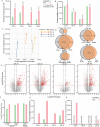
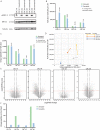
Results: We identified an average of 9936 unique peptides per PDO which compares favorably against published immunopeptidomics studies, suggesting high sensitivity. Loss of heterozygosity of the HLA locus was associated with low peptide diversity in one PDO. Peptides from genes without detectable expression by RNA-sequencing were rarely identified by MS. Only 3 out of 612 non-silent mutations encoded for neoantigens that were detected by MS. In contrast, computational HLA binding prediction estimated that 304 mutations could generate neoantigens. One hundred ninety-six of these were located in expressed genes, still exceeding the number of MS-detected neoantigens 65-fold. Treatment of four PDOs with IFNγ upregulated HLA class I expression and qualitatively changed the immunopeptidome, with increased presentation of IFNγ-inducible genes. HLA class II presented peptides increased dramatically with IFNγ treatment. MEK-inhibitor treatment showed no consistent effect on HLA class I or II expression or the peptidome. Importantly, no additional HLA class I or II presented neoantigens became detectable with any treatment.
Conclusions: Only 3 out of 612 non-silent mutations encoded for neoantigens that were detectable by MS. Although MS has sensitivity limits and biases, and likely underestimated the true neoantigen burden, this established a lower bound of the percentage of non-silent mutations that encode for presented neoantigens, which may be as low as 0.5%. This could be a reason for the poor responses of non-hypermutated CRCs to immune checkpoint inhibitors. MEK-inhibitors recently failed to improve checkpoint-inhibitor efficacy in CRC and the observed lack of HLA upregulation or improved peptide presentation may explain this.
Keywords: Antigen presentation; Colorectal cancer; Human leukocyte antigen; Immunogenicity; Immunotherapy; Mass spectrometry; Neoantigens; Patient derived organoids.
Conflict of interest statement
The authors have no relevant competing interests to declare.
Figures

References
-
- De Weger VA, Turksma AW, Voorham QJM, Euler Z, Bril H, Van Den Eertwegh AJ, et al. Clinical effects of adjuvant active specific immunotherapy differ between patients with microsatellite-stable and microsatellite-instable colon cancer. Clin Cancer Res. 2012;18(3):882–889. doi: 10.1158/1078-0432.CCR-11-1716. - DOI - PubMed
-
- Grasso CS, Giannakis M, Wells DK, Hamada T, Mu XJ, Quist M, et al. Genetic mechanisms of immune evasion in colorectal cancer. Cancer Discov. 2018;8(6):730–749. doi: 10.1158/2159-8290.CD-17-1327. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
