Structures of 1:1 and 2:1 complexes of BMVC and MYC promoter G-quadruplex reveal a mechanism of ligand conformation adjustment for G4-recognition
- PMID: 31740959
- PMCID: PMC7145684
- DOI: 10.1093/nar/gkz1015
Structures of 1:1 and 2:1 complexes of BMVC and MYC promoter G-quadruplex reveal a mechanism of ligand conformation adjustment for G4-recognition
Abstract
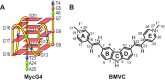
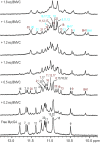
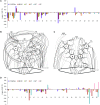
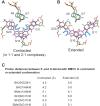
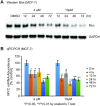
BMVC is the first fluorescent probe designed to detect G-quadruplexes (G4s) in vivo. The MYC oncogene promoter forms a G4 (MycG4) which acts as a transcription silencer. Here, we report the high-affinity and specific binding of BMVC to MycG4 with unusual slow-exchange rates on the NMR timescale. We also show that BMVC represses MYC in cancer cells. We determined the solution structures of the 1:1 and 2:1 BMVC-MycG4 complexes. BMVC first binds the 5'-end of MycG4 to form a 1:1 complex with a well-defined structure. At higher ratio, BMVC also binds the 3'-end to form a second complex. In both complexes, the crescent-shaped BMVC recruits a flanking DNA residue to form a BMVC-base plane stacking over the external G-tetrad. Remarkably, BMVC adjusts its conformation to a contracted form to match the G-tetrad for an optimal stacking interaction. This is the first structural example showing the importance of ligand conformational adjustment in G4 recognition. BMVC binds the more accessible 5'-end with higher affinity, whereas sequence specificity is present at the weaker-binding 3'-site. Our structures provide insights into specific recognition of MycG4 by BMVC and useful information for design of G4-targeted anticancer drugs and fluorescent probes.
© The Author(s) 2019. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures



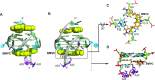
References
-
- Neidle S., Parkinson G.. Telomere maintenance as a target for anticancer drug discovery. Nat. Rev. Drug Discov. 2002; 1:383–393. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

