Up-regulation of cryptochrome 1 gene expression in cotton bollworm (Helicoverpa armigera) during migration over the Bohai Sea
- PMID: 31741806
- PMCID: PMC6859876
- DOI: 10.7717/peerj.8071
Up-regulation of cryptochrome 1 gene expression in cotton bollworm (Helicoverpa armigera) during migration over the Bohai Sea
Abstract
Cryptochromes (CRYs) are flavoproteins and play a pivotal role in circadian clocks which mediate behavior of organisms such as feeding, mating and migrating navigation. Herein, we identified novel transcripts in Helicoverpa armigera of six isoforms of cry1 and seven isoforms of cry2 by Sanger sequencing. Phylogenetic analysis showed that the transcripts of cry1 and cry2 align closely with other insect crys, indicating within-species divergence of Hacry. A dn/ds analysis revealed that the encoding sequence of the cry1 was under purifying selection by a strong negative selection pressure whereas the cry2 was less constraint and showed a less strong purification selection than cry1. In general, Hacrys were more abundantly transcribed in wild migrating populations than that in laboratory maintained populations, and expression of the cry2 was lower than cry1 in all samples tested. Moreover, when compared with the migrating parental population, offspring reared in laboratory conditions showed a significant reduction on transcription of the cry1 but not cry2. These results strongly suggest that cry1 was more related to the migration behavior of H. armigera than cry2.
Keywords: Cryptochromes; Expression analysis; Helicoverpa armigera; Migration; Transcripts.
© 2019 Yang et al.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


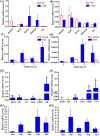
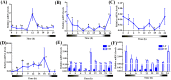
References
-
- Baik LS, Fogle KJ, Roberts L, Galschiodt AM, Chevez JA, Recinos Y, Nguy V, Holmes TC. CRYPTOCHROME mediates behavioral executive choice in response to UV light. Proceedings of the National Academy of Sciences of the United States of America. 2017;114(4):776–781. doi: 10.1073/pnas.1607989114. - DOI - PMC - PubMed
-
- Bazalova O, Kvicalova M, Valkova T, Slaby P, Bartos P, Netusil R, Tomanova K, Braeunig P, Leee H, Saumana I, Damulewicz M, Provaznik J, Pokorny R, Dolezel D, Vacha M. Cryptochrome 2 mediates directional magnetoreception in cockroaches. Proceedings of the National Academy of Sciences of the United States of America. 2016;113(6):1660–1665. doi: 10.1073/pnas.1518622113. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources

