HNF4 Regulates Fatty Acid Oxidation and Is Required for Renewal of Intestinal Stem Cells in Mice
- PMID: 31759926
- PMCID: PMC7062567
- DOI: 10.1053/j.gastro.2019.11.031
HNF4 Regulates Fatty Acid Oxidation and Is Required for Renewal of Intestinal Stem Cells in Mice
Abstract
Background & aims: Functions of intestinal stem cells (ISCs) are regulated by diet and metabolic pathways. Hepatocyte nuclear factor 4 (HNF4) family are transcription factors that bind fatty acids. We investigated how HNF4 transcription factors regulate metabolism and their functions in ISCs in mice.
Methods: We performed studies with Villin-CreERT2;Lgr5-EGFP-IRES-CreERT2;Hnf4αf/f;Hnf4γCrispr/Crispr mice, hereafter referred to Hnf4αγDKO. Mice were given tamoxifen to induce Cre recombinase. Mice transgenic with only Cre alleles (Villin-CreERT2, Lgr5-EGFP-IRES-CreERT2, Hnf4α+/+, and Hnf4γ+/+) or mice given vehicle were used as controls. Crypt and villus cells were isolated, incubated with fluorescently labeled fatty acids or glucose analog, and analyzed by confocal microscopy. Fatty acid oxidation activity and tricarboxylic acid (TCA) cycle metabolites were measured in cells collected from the proximal half of the small intestine of Hnf4αγDKO and control mice. We performed chromatin immunoprecipitation and gene expression profiling analyses to identify genes regulated by HNF4 factors. We established organoids from duodenal crypts, incubated them with labeled palmitate or acetate, and measured production of TCA cycle metabolites or fatty acids. Acetate, a precursor of acetyl coenzyme A (CoA) (a product of fatty acid β-oxidation [FAO]), or dichloroacetate, a compound that promotes pyruvate oxidation and generation of mitochondrial acetyl-CoA, were used for metabolic intervention.
Results: Crypt cells rapidly absorbed labeled fatty acids, and messenger RNA levels of Lgr5+ stem cell markers (Lgr5, Olfm4, Smoc2, Msi1, and Ascl2) were down-regulated in organoids incubated with etomoxir, an inhibitor of FAO, indicating that FAO was required for renewal of ISCs. HNF4A and HNF4G were expressed in ISCs and throughout the intestinal epithelium. Single knockout of either HNF4A or HNF4G did not affect maintenance of ISCs, but double-knockout of HNF4A and HNF4G resulted in ISC loss; stem cells failed to renew. FAO supports ISC renewal, and HNF4 transcription factors directly activate FAO genes, including Acsl5 and Acsf2 (encode regulators of acyl-CoA synthesis), Slc27a2 (encodes a fatty acid transporter), Fabp2 (encodes fatty acid binding protein), and Hadh (encodes hydroxyacyl-CoA dehydrogenase). In the intestinal epithelium of Hnf4αγDKO mice, expression levels of FAO genes, FAO activity, and metabolites of TCA cycle were all significantly decreased, but fatty acid synthesis transcripts were increased, compared with control mice. The contribution of labeled palmitate or acetate to the TCA cycle was reduced in organoids derived from Hnf4αγDKO mice, compared with control mice. Incubation of organoids derived from double-knockout mice with acetate or dichloroacetate restored stem cells.
Conclusions: In mice, the transcription factors HNF4A and HNF4G regulate the expression of genes required for FAO and are required for renewal of ISCs.
Keywords: Gene Expression; Mitochondria; Regulatory Networks; Stemness.
Copyright © 2020 AGA Institute. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflicts of interest
The authors declare no competing interests.
Figures
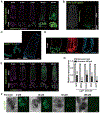




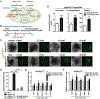
References
-
- Barker N, van Es JH, Kuipers J, et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 2007;449:1003–7. - PubMed
-
- Barker N, Ridgway RA, van Es JH, et al. Crypt stem cells as the cells-of-origin of intestinal cancer. Nature 2009;457:608–11. - PubMed
-
- Metcalfe C, Kljavin NM, Ybarra R, et al. Lgr5+ stem cells are indispensable for radiation-induced intestinal regeneration. Cell Stem Cell 2014;14:149–59. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

