Development and optimization of a hybridization technique to type the classical class I and class II B genes of the chicken MHC
- PMID: 31761978
- PMCID: PMC6900278
- DOI: 10.1007/s00251-019-01149-2
Development and optimization of a hybridization technique to type the classical class I and class II B genes of the chicken MHC
Abstract
The classical class I and class II molecules of the major histocompatibility complex (MHC) play crucial roles in immune responses to infectious pathogens and vaccines as well as being important for autoimmunity, allergy, cancer and reproduction. These classical MHC genes are the most polymorphic known, with roughly 10,000 alleles in humans. In chickens, the MHC (also known as the BF-BL region) determines decisive resistance and susceptibility to infectious pathogens, but relatively few MHC alleles and haplotypes have been described in any detail. We describe a typing protocol for classical chicken class I (BF) and class II B (BLB) genes based on a hybridization method called reference strand-mediated conformational analysis (RSCA). We optimize the various steps, validate the analysis using well-characterized chicken MHC haplotypes, apply the system to type some experimental lines and discover a new chicken class I allele. This work establishes a basis for typing the MHC genes of chickens worldwide and provides an opportunity to correlate with microsatellite and with single nucleotide polymorphism (SNP) typing for approaches involving imputation.
Keywords: Avian; BF1; BF2; BLB1; BLB2; Recombination.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


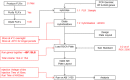
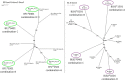


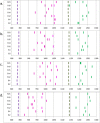

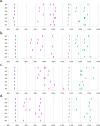
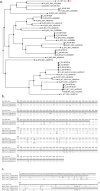
References
-
- Argüello JR, Little AM, Bohan E, Goldman JM, Marsh SG, Madrigal JA. High resolution HLA class I typing by reference strand mediated conformation analysis (RSCA) Tissue Antigens. 1998;52:57–66. - PubMed
-
- Atkinson D, Shaw I, Jacob J, Kaufman J (2001) DM gene polymorphisms: co-evolution or coincidence? In Proceedings of the Avian Immunology Research Group, 7-10 October 2000. Ithaca NY Edited by KA Schat:163–165
-
- Babik W. Methods for MHC typing in non-model vertebrates. Mol Ecol Resour. 2010;10:237–251. - PubMed
-
- Boonyanuwat K, Thummabutra S, Sookmanee N, Vatchavalkhu V, Siripholvat V. Influences of major histocompatibility complex class I haplotypes on avian influenza virus disease traits in Thai indigenous chickens. Anim Sci J. 2006;77:285–289.
-
- Bryceson YT, Chiang SC, Darmanin S, Fauriat C, Schlums H, Theorell J, Wood SM. Molecular mechanisms of natural killer cell activation. J Innate Immun. 2011;3:216–226. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

