Bottom-up structural proteomics: cryoEM of protein complexes enriched from the cellular milieu
- PMID: 31768063
- PMCID: PMC7494424
- DOI: 10.1038/s41592-019-0637-y
Bottom-up structural proteomics: cryoEM of protein complexes enriched from the cellular milieu
Abstract
X-ray crystallography often requires non-native constructs involving mutations or truncations, and is challenged by membrane proteins and large multicomponent complexes. We present here a bottom-up endogenous structural proteomics approach whereby near-atomic-resolution cryo electron microscopy (cryoEM) maps are reconstructed ab initio from unidentified protein complexes enriched directly from the endogenous cellular milieu, followed by identification and atomic modeling of the proteins. The proteins in each complex are identified using cryoID, a program we developed to identify proteins in ab initio cryoEM maps. As a proof of principle, we applied this approach to the malaria-causing parasite Plasmodium falciparum, an organism that has resisted conventional structural-biology approaches, to obtain atomic models of multiple protein complexes implicated in intraerythrocytic survival of the parasite. Our approach is broadly applicable for determining structures of undiscovered protein complexes enriched directly from endogenous sources.
Figures

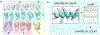



References
-
- Clough GMRN, Kirkland AI. in Journal of Physics: Conference Series Vol. 522 (IOP Publishing, 2013).
Publication types
MeSH terms
Substances
Grants and funding
- R00 HL133453/HL/NHLBI NIH HHS/United States
- S10OD018111/U.S. Department of Health & Human Services | NIH | NIH Office of the Director (OD)/International
- R01 AI094386/AI/NIAID NIH HHS/United States
- U24GM116792/U.S. Department of Health & Human Services | NIH | National Institute of General Medical Sciences (NIGMS)/International
- S10 OD018111/OD/NIH HHS/United States
- S10RR23057/U.S. Department of Health & Human Services | NIH | National Institute of Dental and Craniofacial Research (NIDCR)/International
- S10 RR023057/RR/NCRR NIH HHS/United States
- R01 GM089778/GM/NIGMS NIH HHS/United States
- T32 AI007323/AI/NIAID NIH HHS/United States
- R01 GM112763/GM/NIGMS NIH HHS/United States
- K99/R00 HL133453/U.S. Department of Health & Human Services | NIH | National Heart, Lung, and Blood Institute (NHLBI)/International
- K99 HL133453/HL/NHLBI NIH HHS/United States
- U24 GM116792/GM/NIGMS NIH HHS/United States
- R01 GM071940/GM/NIGMS NIH HHS/United States
- R01GM071940/U.S. Department of Health & Human Services | NIH | National Institute of General Medical Sciences (NIGMS)/International
- R01 DE025567/DE/NIDCR NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

