Eukaryote hybrid genomes
- PMID: 31774811
- PMCID: PMC6880984
- DOI: 10.1371/journal.pgen.1008404
Eukaryote hybrid genomes
Abstract
Interspecific hybridization is the process where closely related species mate and produce offspring with admixed genomes. The genomic revolution has shown that hybridization is common, and that it may represent an important source of novel variation. Although most interspecific hybrids are sterile or less fit than their parents, some may survive and reproduce, enabling the transfer of adaptive variants across the species boundary, and even result in the formation of novel evolutionary lineages. There are two main variants of hybrid species genomes: allopolyploid, which have one full chromosome set from each parent species, and homoploid, which are a mosaic of the parent species genomes with no increase in chromosome number. The establishment of hybrid species requires the development of reproductive isolation against parental species. Allopolyploid species often have strong intrinsic reproductive barriers due to differences in chromosome number, and homoploid hybrids can become reproductively isolated from the parent species through assortment of genetic incompatibilities. However, both types of hybrids can become further reproductively isolated, gaining extrinsic isolation barriers, by exploiting novel ecological niches, relative to their parents. Hybrids represent the merging of divergent genomes and thus face problems arising from incompatible combinations of genes. Thus hybrid genomes are highly dynamic and undergo rapid evolutionary change, including genome stabilization in which selection against incompatible combinations results in fixation of compatible ancestry block combinations within the hybrid species. The potential for rapid adaptation or speciation makes hybrid genomes a particularly exciting subject of in evolutionary biology. Here we summarize how introgressed alleles or hybrid species can establish and how the resulting hybrid genomes evolve.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
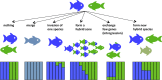


References
-
- Abbott R, Albach D, Ansell S, Arntzen J, Baird S, Bierne N, et al. (2013), "Hybridization and speciation", Journal of Evolutionary Biology 26 (2): 229–246, 10.1111/j.1420-9101.2012.02599.x http://doi.wiley.com/10.1111/j.1420-9101.2012.02599.x - DOI - DOI - PubMed
-
- Fisher R (1930), The genetical theory of natural selection., Oxford: Clarendon Press, 10.5962/bhl.title.27468 http://www.biodiversitylibrary.org/bibliography/27468 - DOI
-
- Mayr E (1963), Animal Species and Evolution:, Cambridge, MA and London, England: Harvard University Press, 10.4159/harvard.9780674865327 ISBN 9780674865327, http://www.degruyter.com/view/books/harvard.9780674865327/harvard.978067... - DOI
-
- Stebbins G (1959), "The Role of Hybridization in Evolution", Proceedings of the American Philosophical Society 103 (2): 231–251, ISSN 0003-049X, https://www.jstor.org/stable/985151
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

