Serratia liquefaciens FG3 isolated from a metallophyte plant sheds light on the evolution and mechanisms of adaptive traits in extreme environments
- PMID: 31784663
- PMCID: PMC6884506
- DOI: 10.1038/s41598-019-54601-4
Serratia liquefaciens FG3 isolated from a metallophyte plant sheds light on the evolution and mechanisms of adaptive traits in extreme environments
Abstract
Serratia liquefaciens strain FG3 (SlFG3), isolated from the flower of Stachytarpheta glabra in the Brazilian ferruginous fields, has distinctive genomic, adaptive, and biotechnological potential. Herein, using a combination of genomics and molecular approaches, we unlocked the evolution of the adaptive traits acquired by S1FG3, which exhibits the second largest chromosome containing the largest conjugative plasmids described for Serratia. Comparative analysis revealed the presence of 18 genomic islands and 311 unique protein families involved in distinct adaptive features. S1FG3 has a diversified repertoire of genes associated with Nonribosomal peptides (NRPs/PKS), a complete and functional cluster related to cellulose synthesis, and an extensive and functional repertoire of oxidative metabolism genes. In addition, S1FG3 possesses a complete pathway related to protocatecuate and chloroaromatic degradation, and a complete repertoire of genes related to DNA repair and protection that includes mechanisms related to UV light tolerance, redox process resistance, and a laterally acquired capacity to protect DNA using phosphorothioation. These findings summarize that SlFG3 is well-adapted to different biotic and abiotic stress situations imposed by extreme conditions associated with ferruginous fields, unlocking the impact of the lateral gene transfer to adjust the genome for extreme environments, and providing insight into the evolution of prokaryotes.
Conflict of interest statement
The authors declare no competing interests.
Figures

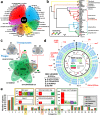






References
-
- Sandner-Miranda, L., Vinuesa, P., Soberon-Chavez, G. & Morales-Espinosa, R. Complete Genome Sequence of Serratia marcescens SmUNAM836, a Nonpigmented Multidrug-Resistant Strain Isolated from a Mexican Patient with Obstructive Pulmonary Disease. Genome announcements, 4, 10.1128/genomeA.01417-15 (2016). - PMC - PubMed
-
- Adam E, Muller H, Erlacher A, Berg G. Complete genome sequences of the Serratia plymuthica strains 3Rp8 and 3Re4-18, two rhizosphere bacteria with antagonistic activity towards fungal phytopathogens and plant growth promoting abilities. Standards in genomic sciences. 2016;11:61. doi: 10.1186/s40793-016-0185-3. - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

