A cancer rainbow mouse for visualizing the functional genomics of oncogenic clonal expansion
- PMID: 31792216
- PMCID: PMC6889384
- DOI: 10.1038/s41467-019-13330-y
A cancer rainbow mouse for visualizing the functional genomics of oncogenic clonal expansion
Abstract
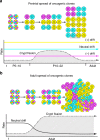
Field cancerization is a premalignant process marked by clones of oncogenic mutations spreading through the epithelium. The timescales of intestinal field cancerization can be variable and the mechanisms driving the rapid spread of oncogenic clones are unknown. Here we use a Cancer rainbow (Crainbow) modelling system for fluorescently barcoding somatic mutations and directly visualizing the clonal expansion and spread of oncogenes. Crainbow shows that mutations of ß-catenin (Ctnnb1) within the intestinal stem cell results in widespread expansion of oncogenes during perinatal development but not in adults. In contrast, mutations that extrinsically disrupt the stem cell microenvironment can spread in adult intestine without delay. We observe the rapid spread of premalignant clones in Crainbow mice expressing oncogenic Rspondin-3 (RSPO3), which occurs by increasing crypt fission and inhibiting crypt fixation. Crainbow modelling provides insight into how somatic mutations rapidly spread and a plausible mechanism for predetermining the intratumor heterogeneity found in colon cancers.
Conflict of interest statement
The authors declare no competing interests.
Figures







References
-
- Braakhuis BJ, Tabor MP, Kummer JA, Leemans CR, Brakenhoff RH. A genetic explanation of Slaughter’s concept of field cancerization: evidence and clinical implications. Cancer Res. 2003;63:1727–1730. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

