Molecular replacement using structure predictions from databases
- PMID: 31793899
- PMCID: PMC6889911
- DOI: 10.1107/S2059798319013962
Molecular replacement using structure predictions from databases
Abstract
Molecular replacement (MR) is the predominant route to solution of the phase problem in macromolecular crystallography. Where the lack of a suitable homologue precludes conventional MR, one option is to predict the target structure using bioinformatics. Such modelling, in the absence of homologous templates, is called ab initio or de novo modelling. Recently, the accuracy of such models has improved significantly as a result of the availability, in many cases, of residue-contact predictions derived from evolutionary covariance analysis. Covariance-assisted ab initio models representing structurally uncharacterized Pfam families are now available on a large scale in databases, potentially representing a valuable and easily accessible supplement to the PDB as a source of search models. Here, the unconventional MR pipeline AMPLE is employed to explore the value of structure predictions in the GREMLIN and PconsFam databases. It was tested whether these deposited predictions, processed in various ways, could solve the structures of PDB entries that were subsequently deposited. The results were encouraging: nine of 27 GREMLIN cases were solved, covering target lengths of 109-355 residues and a resolution range of 1.4-2.9 Å, and with target-model shared sequence identity as low as 20%. The cluster-and-truncate approach in AMPLE proved to be essential for most successes. For the overall lower quality structure predictions in the PconsFam database, remodelling with Rosetta within the AMPLE pipeline proved to be the best approach, generating ensemble search models from single-structure deposits. Finally, it is shown that the AMPLE-obtained search models deriving from GREMLIN deposits are of sufficiently high quality to be selected by the sequence-independent MR pipeline SIMBAD. Overall, the results help to point the way towards the optimal use of the expanding databases of ab initio structure predictions.
Keywords: ab initio modelling; ab initio structure predictions; databases; molecular replacement.
open access.
Figures
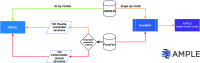



References
-
- Adams, P. D., Afonine, P. V., Bunkóczi, G., Chen, V. B., Davis, I. W., Echols, N., Headd, J. J., Hung, L.-W., Kapral, G. J., Grosse-Kunstleve, R. W., McCoy, A. J., Moriarty, N. W., Oeffner, R., Read, R. J., Richardson, D. C., Richardson, J. S., Terwilliger, T. C. & Zwart, P. H. (2010). Acta Cryst. D66, 213–221. - PMC - PubMed
-
- Alford, R. F., Leaver-Fay, A., Jeliazkov, J. R., O’Meara, M. J., DiMaio, F. P., Park, H., Shapovalov, M. V., Renfrew, P. D., Mulligan, V. K., Kappel, K., Labonte, J. W., Pacella, M. S., Bonneau, R., Bradley, P., Dunbrack, R. L. Jr, Das, R., Baker, D., Kuhlman, B., Kortemme, T. & Gray, J. J. (2017). J. Chem. Theory Comput. 13, 3031–3048. - PMC - PubMed
-
- Bibby, J., Keegan, R. M., Mayans, O., Winn, M. D. & Rigden, D. J. (2012). Acta Cryst. D68, 1622–1631. - PubMed
-
- Brunger, A. T. (2007). Nature Protoc. 2, 2728–2733. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

