Deficiencies in vesicular transport mediated by TRAPPC4 are associated with severe syndromic intellectual disability
- PMID: 31794024
- PMCID: PMC6935753
- DOI: 10.1093/brain/awz374
Deficiencies in vesicular transport mediated by TRAPPC4 are associated with severe syndromic intellectual disability
Erratum in
-
Erratum.Brain. 2020 Mar 1;143(3):e24. doi: 10.1093/brain/awaa007. Brain. 2020. PMID: 32333675 Free PMC article. No abstract available.
Abstract
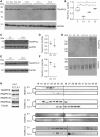
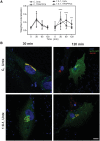
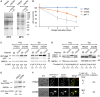
The conserved transport protein particle (TRAPP) complexes regulate key trafficking events and are required for autophagy. TRAPPC4, like its yeast Trs23 orthologue, is a core component of the TRAPP complexes and one of the essential subunits for guanine nucleotide exchange factor activity for Rab1 GTPase. Pathogenic variants in specific TRAPP subunits are associated with neurological disorders. We undertook exome sequencing in three unrelated families of Caucasian, Turkish and French-Canadian ethnicities with seven affected children that showed features of early-onset seizures, developmental delay, microcephaly, sensorineural deafness, spastic quadriparesis and progressive cortical and cerebellar atrophy in an effort to determine the genetic aetiology underlying neurodevelopmental disorders. All seven affected subjects shared the same identical rare, homozygous, potentially pathogenic variant in a non-canonical, well-conserved splice site within TRAPPC4 (hg19:chr11:g.118890966A>G; TRAPPC4: NM_016146.5; c.454+3A>G). Single nucleotide polymorphism array analysis revealed there was no haplotype shared between the tested Turkish and Caucasian families suggestive of a variant hotspot region rather than a founder effect. In silico analysis predicted the variant to cause aberrant splicing. Consistent with this, experimental evidence showed both a reduction in full-length transcript levels and an increase in levels of a shorter transcript missing exon 3, suggestive of an incompletely penetrant splice defect. TRAPPC4 protein levels were significantly reduced whilst levels of other TRAPP complex subunits remained unaffected. Native polyacrylamide gel electrophoresis and size exclusion chromatography demonstrated a defect in TRAPP complex assembly and/or stability. Intracellular trafficking through the Golgi using the marker protein VSVG-GFP-ts045 demonstrated significantly delayed entry into and exit from the Golgi in fibroblasts derived from one of the affected subjects. Lentiviral expression of wild-type TRAPPC4 in these fibroblasts restored trafficking, suggesting that the trafficking defect was due to reduced TRAPPC4 levels. Consistent with the recent association of the TRAPP complex with autophagy, we found that the fibroblasts had a basal autophagy defect and a delay in autophagic flux, possibly due to unsealed autophagosomes. These results were validated using a yeast trs23 temperature sensitive variant that exhibits constitutive and stress-induced autophagic defects at permissive temperature and a secretory defect at restrictive temperature. In summary we provide strong evidence for pathogenicity of this variant in a member of the core TRAPP subunit, TRAPPC4 that associates with vesicular trafficking and autophagy defects. This is the first report of a TRAPPC4 variant, and our findings add to the growing number of TRAPP-associated neurological disorders.
Keywords: autophagy; intellectual disability; molecular genetics; vesicular transport; whole-exome sequencing.
© The Author(s) (2019). Published by Oxford University Press on behalf of the Guarantors of Brain. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Figures




Comment in
-
Reply: Recurrent bi-allelic splicing variant c.454+3A>G in TRAPPC4 is associated with progressive encephalopathy and muscle involvement.Brain. 2020 Apr 1;143(4):e30. doi: 10.1093/brain/awaa047. Brain. 2020. PMID: 32125358 No abstract available.
-
Recurrent bi-allelic splicing variant c.454+3A>G in TRAPPC4 is associated with progressive encephalopathy and muscle involvement.Brain. 2020 Apr 1;143(4):e29. doi: 10.1093/brain/awaa046. Brain. 2020. PMID: 32125366 No abstract available.
-
Biallelic in-frame deletion in TRAPPC4 in a family with developmental delay and cerebellar atrophy.Brain. 2020 Oct 1;143(10):e83. doi: 10.1093/brain/awaa256. Brain. 2020. PMID: 33011761 Free PMC article. No abstract available.
-
Reply: Biallelic in-frame deletion in TRAPPC4 in a family with developmental delay and cerebellar atrophy.Brain. 2020 Oct 1;143(10):e84. doi: 10.1093/brain/awaa257. Brain. 2020. PMID: 33011764 No abstract available.
References
-
- Blomen VA, Majek P, Jae LT, Bigenzahn JW, Nieuwenhuis J, Staring J, et al. Gene essentiality and synthetic lethality in haploid human cells. Science 2015; 350: 1092–6. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

