Exposing Hidden High-Affinity RNA Conformational States
- PMID: 31815464
- PMCID: PMC7380322
- DOI: 10.1021/jacs.9b10535
Exposing Hidden High-Affinity RNA Conformational States
Abstract
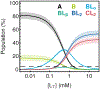
RNA recognition frequently results in conformational changes that optimize intermolecular binding. As a consequence, the overall binding affinity of RNA to its binding partners depends not only on the intermolecular interactions formed in the bound state but also on the energy cost associated with changing the RNA conformational distribution. Measuring these "conformational penalties" is, however, challenging because bound RNA conformations tend to have equilibrium populations in the absence of the binding partner that fall outside detection by conventional biophysical methods. In this study we employ as a model system HIV-1 TAR RNA and its interaction with the ligand argininamide (ARG), a mimic of TAR's cognate protein binding partner, the transactivator Tat. We use NMR chemical shift perturbations and relaxation dispersion in combination with Bayesian inference to develop a detailed thermodynamic model of coupled conformational change and ligand binding. Starting from a comprehensive 12-state model of the equilibrium, we estimate the energies of six distinct detectable thermodynamic states that are not accessible by currently available methods. Our approach identifies a minimum of four RNA intermediates that differ in terms of the TAR conformation and ARG occupancy. The dominant bound TAR conformation features two bound ARG ligands and has an equilibrium population in the absence of ARG that is below detection limit. Consequently, even though ARG binds to TAR with an apparent overall weak affinity (Kdapp ≈ 0.2 mM), it binds the prefolded conformation with a Kd in the nM range. Our results show that conformational penalties can be major determinants of RNA-ligand binding affinity as well as a source of binding cooperativity, with important implications for a predictive understanding of how RNA is recognized and for RNA-targeted drug discovery.
Conflict of interest statement
The authors declare the following competing financial interest(s): H.M.A. is an advisor and holds an ownership interest in Nymirum Inc., an RNA-based drug discovery company.
Figures
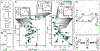




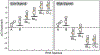
References
-
- Mattick JS RNA regulation: a new genetics? Nat. Rev. Genet 2004, 5, 316–23. - PubMed
-
- Sharp PA The centrality of RNA. Cell 2009, 136, 577–80. - PubMed
-
- Cruz JA; Westhof E The dynamic landscapes of RNA architecture. Cell 2009, 136, 604–9. - PubMed
-
- Rinnenthal J; Buck J; Ferner J; Wacker A; Fürtig B; Schwalbe H Mapping the landscape of RNA dynamics with NMR spectroscopy. Acc. Chem. Res 2011, 44, 1292–1301. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

