Defining a metabolic landscape of tumours: genome meets metabolism
- PMID: 31819196
- PMCID: PMC7051970
- DOI: 10.1038/s41416-019-0663-7
Defining a metabolic landscape of tumours: genome meets metabolism
Abstract
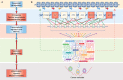
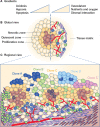
Cancer is a complex disease of multiple alterations occuring at the epigenomic, genomic, transcriptomic, proteomic and/or metabolic levels. The contribution of genetic mutations in cancer initiation, progression and evolution is well understood. However, although metabolic changes in cancer have long been acknowledged and considered a plausible therapeutic target, the crosstalk between genetic and metabolic alterations throughout cancer types is not clearly defined. In this review, we summarise the present understanding of the interactions between genetic drivers of cellular transformation and cancer-associated metabolic changes, and how these interactions contribute to metabolic heterogeneity of tumours. We discuss the essential question of whether changes in metabolism are a cause or a consequence in the formation of cancer. We highlight two modes of how metabolism contributes to tumour formation. One is when metabolic reprogramming occurs downstream of oncogenic mutations in signalling pathways and supports tumorigenesis. The other is where metabolic reprogramming initiates transformation being either downstream of mutations in oncometabolite genes or induced by chronic wounding, inflammation, oxygen stress or metabolic diseases. Finally, we focus on the factors that can contribute to metabolic heterogeneity in tumours, including genetic heterogeneity, immunomodulatory factors and tissue architecture. We believe that an in-depth understanding of cancer metabolic reprogramming, and the role of metabolic dysregulation in tumour initiation and progression, can help identify cellular vulnerabilities that can be exploited for therapeutic use.
Conflict of interest statement
The authors declare no competing interests.
Figures
References
-
- Plaxton NA, Brandon DC, Corey AS, Harrison CE, Tuba Karagulle Kendi A, Halkar RK, et al. Characteristics and Limitations of FDG PET/CT for imaging of squamous cell carcinoma of the head and neck: a comprehensive review of anatomy, metastatic pathways, and image findings. Am. J. Roentgenol. 2015;205:W519–W531. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

