Loss of IL-10 signaling in macrophages limits bacterial killing driven by prostaglandin E2
- PMID: 31819956
- PMCID: PMC7041704
- DOI: 10.1084/jem.20180649
Loss of IL-10 signaling in macrophages limits bacterial killing driven by prostaglandin E2
Abstract
Loss of IL-10 signaling in macrophages (Mφs) leads to inflammatory bowel disease (IBD). Induced pluripotent stem cells (iPSCs) were generated from an infantile-onset IBD patient lacking a functional IL10RB gene. Mφs differentiated from IL-10RB-/- iPSCs lacked IL-10RB mRNA expression, were unable to phosphorylate STAT3, and failed to reduce LPS induced inflammatory cytokines in the presence of exogenous IL-10. IL-10RB-/- Mφs exhibited a striking defect in their ability to kill Salmonella enterica serovar Typhimurium, which was rescuable after experimentally introducing functional copies of the IL10RB gene. Genes involved in synthesis and receptor pathways for eicosanoid prostaglandin E2 (PGE2) were more highly induced in IL-10RB-/- Mφs, and these Mφs produced higher amounts of PGE2 after LPS stimulation compared with controls. Furthermore, pharmacological inhibition of PGE2 synthesis and PGE2 receptor blockade enhanced bacterial killing in Mφs. These results identify a regulatory interaction between IL-10 and PGE2, dysregulation of which may drive aberrant Mφ activation and impaired host defense contributing to IBD pathogenesis.
© 2019 Mukhopadhyay et al.
Figures

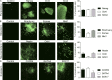




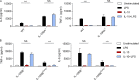
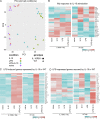




Similar articles
-
Genetic association and epistatic interaction of the interleukin-10 signaling pathway in pediatric inflammatory bowel disease.World J Gastroenterol. 2017 Jul 21;23(27):4897-4909. doi: 10.3748/wjg.v23.i27.4897. World J Gastroenterol. 2017. PMID: 28785144 Free PMC article.
-
Knockout-Induced Pluripotent Stem Cells for Disease and Therapy Modeling of IL-10-Associated Primary Immunodeficiencies.Hum Gene Ther. 2021 Jan;32(1-2):77-95. doi: 10.1089/hum.2020.235. Epub 2021 Jan 11. Hum Gene Ther. 2021. PMID: 33023317
-
[Interleukin-10 receptor mutations in children with neonatal onset inflammatory bowel disease: genetic diagnosis and pathogenesis].Zhonghua Er Ke Za Zhi. 2015 May;53(5):348-54. Zhonghua Er Ke Za Zhi. 2015. PMID: 26080664 Chinese.
-
The Tpl2 Kinase Regulates the COX-2/Prostaglandin E2 Axis in Adipocytes in Inflammatory Conditions.Mol Endocrinol. 2015 Jul;29(7):1025-36. doi: 10.1210/me.2015-1027. Epub 2015 May 28. Mol Endocrinol. 2015. PMID: 26020725 Free PMC article.
-
Very early-onset inflammatory bowel disease (IBD) in infancy is a different disease entity from adult-onset IBD; one form of interleukin-10 receptor mutations.J Hum Genet. 2014 Jun;59(6):337-41. doi: 10.1038/jhg.2014.32. Epub 2014 May 1. J Hum Genet. 2014. PMID: 24785691
Cited by
-
IL-23 Contributes to Campylobacter jejuni-Induced Intestinal Pathology via Promoting IL-17 and IFNγ Responses by Innate Lymphoid Cells.Front Immunol. 2021 Jan 6;11:579615. doi: 10.3389/fimmu.2020.579615. eCollection 2020. Front Immunol. 2021. PMID: 33488580 Free PMC article.
-
A Multicellular In Vitro Model of the Human Intestine with Immunocompetent Features Highlights Host-Pathogen Interactions During Early Salmonella Typhimurium Infection.Adv Sci (Weinh). 2025 Mar;12(9):e2411233. doi: 10.1002/advs.202411233. Epub 2025 Jan 14. Adv Sci (Weinh). 2025. PMID: 39807570 Free PMC article.
-
Turning foes into permissive hosts: manipulation of macrophage polarization by intracellular bacteria.Curr Opin Immunol. 2023 Oct;84:102367. doi: 10.1016/j.coi.2023.102367. Epub 2023 Jul 10. Curr Opin Immunol. 2023. PMID: 37437470 Free PMC article. Review.
-
Induced Pluripotent Stem Cell-Derived Monocytes/Macrophages in Autoinflammatory Diseases.Front Immunol. 2022 May 6;13:870535. doi: 10.3389/fimmu.2022.870535. eCollection 2022. Front Immunol. 2022. PMID: 35603217 Free PMC article. Review.
-
Deconvolution of monocyte responses in inflammatory bowel disease reveals an IL-1 cytokine network that regulates IL-23 in genetic and acquired IL-10 resistance.Gut. 2021 Jun;70(6):1023-1036. doi: 10.1136/gutjnl-2020-321731. Epub 2020 Oct 9. Gut. 2021. PMID: 33037057 Free PMC article.
References
-
- Agu C.A., Soares F.A., Alderton A., Patel M., Ansari R., Patel S., Forrest S., Yang F., Lineham J., Vallier L., and Kirton C.M.. 2015. Successful generation of human induced pluripotent stem cell lines from blood samples held at room temperature for up to 48 hr. Stem Cell Reports. 5:660–671. 10.1016/j.stemcr.2015.08.012 - DOI - PMC - PubMed
-
- Alasoo K., Martinez F.O., Hale C., Gordon S., Powrie F., Dougan G., Mukhopadhyay S., and Gaffney D.J.. 2015. Transcriptional profiling of macrophages derived from monocytes and iPS cells identifies a conserved response to LPS and novel alternative transcription. Sci. Rep. 5:12524 10.1038/srep12524 - DOI - PMC - PubMed
-
- Arai Y., Arihiro S., Matsuura T., Kato T., Matsuoka M., Saruta M., Mitsunaga M., Matsuura M., Fujiwara M., Okayasu I., et al. . 2014. Prostaglandin E-major urinary metabolite as a reliable surrogate marker for mucosal inflammation in ulcerative colitis. Inflamm. Bowel Dis. 20:1208–1216. 10.1097/MIB.0000000000000062 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

