GenoSurf: metadata driven semantic search system for integrated genomic datasets
- PMID: 31820804
- PMCID: PMC6902006
- DOI: 10.1093/database/baz132
GenoSurf: metadata driven semantic search system for integrated genomic datasets
Abstract
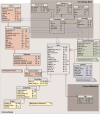
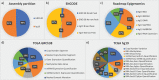
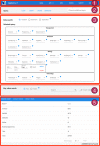
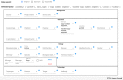
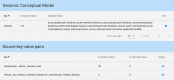
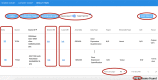
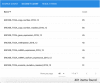
Many valuable resources developed by world-wide research institutions and consortia describe genomic datasets that are both open and available for secondary research, but their metadata search interfaces are heterogeneous, not interoperable and sometimes with very limited capabilities. We implemented GenoSurf, a multi-ontology semantic search system providing access to a consolidated collection of metadata attributes found in the most relevant genomic datasets; values of 10 attributes are semantically enriched by making use of the most suited available ontologies. The user of GenoSurf provides as input the search terms, sets the desired level of ontological enrichment and obtains as output the identity of matching data files at the various sources. Search is facilitated by drop-down lists of matching values; aggregate counts describing resulting files are updated in real time while the search terms are progressively added. In addition to the consolidated attributes, users can perform keyword-based searches on the original (raw) metadata, which are also imported; GenoSurf supports the interplay of attribute-based and keyword-based search through well-defined interfaces. Currently, GenoSurf integrates about 40 million metadata of several major valuable data sources, including three providers of clinical and experimental data (TCGA, ENCODE and Roadmap Epigenomics) and two sources of annotation data (GENCODE and RefSeq); it can be used as a standalone resource for targeting the genomic datasets at their original sources (identified with their accession IDs and URLs), or as part of an integrated query answering system for performing complex queries over genomic regions and metadata.
© The Author(s) 2019. Published by Oxford University Press.
Figures


References
-
- Bernasconi A., Ceri S., Campi A. et al. (2017) Conceptual modeling for genomics: building an integrated repository of open data In: Proceedings of Conceptual Modeling - 36th International Conference (ER 2017). Valencia, Spain, pp. 325–339.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

