ZPR1 prevents R-loop accumulation, upregulates SMN2 expression and rescues spinal muscular atrophy
- PMID: 31828288
- PMCID: PMC6935747
- DOI: 10.1093/brain/awz373
ZPR1 prevents R-loop accumulation, upregulates SMN2 expression and rescues spinal muscular atrophy
Erratum in
-
Erratum.Brain. 2020 Mar 1;143(3):e24. doi: 10.1093/brain/awaa007. Brain. 2020. PMID: 32333675 Free PMC article. No abstract available.
Abstract
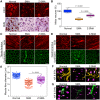
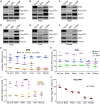
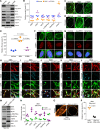
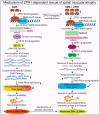
Spinal muscular atrophy (SMA) is a neuromuscular disorder caused by homozygous mutation or deletion of the survival motor neuron 1 (SMN1) gene. A second copy, SMN2, is similar to SMN1 but produces ∼10% SMN protein because of a single-point mutation that causes splicing defects. Chronic low levels of SMN cause accumulation of co-transcriptional R-loops and DNA damage leading to genomic instability and neurodegeneration in SMA. Severity of SMA disease correlates inversely with SMN levels. SMN2 is a promising target to produce higher levels of SMN by enhancing its expression. Mechanisms that regulate expression of SMN genes are largely unknown. We report that zinc finger protein ZPR1 binds to RNA polymerase II, interacts in vivo with SMN locus and upregulates SMN2 expression in SMA mice and patient cells. Modulation of ZPR1 levels directly correlates and influences SMN2 expression levels in SMA patient cells. ZPR1 overexpression in vivo results in a systemic increase of SMN levels and rescues severe to moderate disease in SMA mice. ZPR1-dependent rescue improves growth and motor function and increases the lifespan of male and female SMA mice. ZPR1 reduces neurodegeneration in SMA mice and prevents degeneration of cultured primary spinal cord neurons derived from SMA mice. Further, we show that the low levels of ZPR1 associated with SMA pathogenesis cause accumulation of co-transcriptional RNA-DNA hybrids (R-loops) and DNA damage leading to genomic instability in SMA mice and patient cells. Complementation with ZPR1 elevates senataxin levels, reduces R-loop accumulation and rescues DNA damage in SMA mice, motor neurons and patient cells. In conclusion, ZPR1 is critical for preventing accumulation of co-transcriptional R-loops and DNA damage to avert genomic instability and neurodegeneration in SMA. ZPR1 enhances SMN2 expression and leads to SMN-dependent rescue of SMA. ZPR1 represents a protective modifier and a therapeutic target for developing a new method for the treatment of SMA.
Keywords: R-loops; SMN; ZPR1; neurodegeneration; spinal muscular atrophy.
© The Author(s) (2019). Published by Oxford University Press on behalf of the Guarantors of Brain.
Figures




Comment in
-
Resolution of pathogenic R-loops rescues motor neuron degeneration in spinal muscular atrophy.Brain. 2020 Jan 1;143(1):2-5. doi: 10.1093/brain/awz394. Brain. 2020. PMID: 31886489 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

