Engineering State-of-the-Art Plasmonic Nanomaterials for SERS-Based Clinical Liquid Biopsy Applications
- PMID: 31832306
- PMCID: PMC6891916
- DOI: 10.1002/advs.201900730
Engineering State-of-the-Art Plasmonic Nanomaterials for SERS-Based Clinical Liquid Biopsy Applications
Abstract
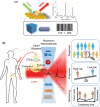
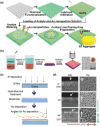
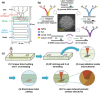
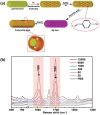
Precision oncology, defined as the use of the molecular understanding of cancer to implement personalized patient treatment, is currently at the heart of revolutionizing oncology practice. Due to the need for repeated molecular tumor analyses in facilitating precision oncology, liquid biopsies, which involve the detection of noninvasive cancer biomarkers in circulation, may be a critical key. Yet, existing liquid biopsy analysis technologies are still undergoing an evolution to address the challenges of analyzing trace quantities of circulating tumor biomarkers reliably and cost effectively. Consequently, the recent emergence of cutting-edge plasmonic nanomaterials represents a paradigm shift in harnessing the unique merits of surface-enhanced Raman scattering (SERS) biosensing platforms for clinical liquid biopsy applications. Herein, an expansive review on the design/synthesis of a new generation of diverse plasmonic nanomaterials, and an updated evaluation of their demonstrated SERS-based uses in liquid biopsies, such as circulating tumor cells, tumor-derived extracellular vesicles, as well as circulating cancer proteins, and tumor nucleic acids is presented. Existing challenges impeding the clinical translation of plasmonic nanomaterials for SERS-based liquid biopsy applications are also identified, and outlooks and insights into advancing this rapidly growing field for practical patient use are provided.
Keywords: clinical translation; liquid biopsies; plasmonic nanomaterials; precision oncology; surface‐enhanced Raman scattering.
© 2019 The Authors. Published by WILEY‐VCH Verlag GmbH & Co. KGaA, Weinheim.
Conflict of interest statement
The authors declare no conflict of interest.
Figures













Similar articles
-
SERS biosensors for liquid biopsy towards cancer diagnosis by detection of various circulating biomarkers: current progress and perspectives.Nano Converg. 2024 May 29;11(1):22. doi: 10.1186/s40580-024-00428-3. Nano Converg. 2024. PMID: 38811455 Free PMC article. Review.
-
Plasmonic substrates for biochemical applications of surface-enhanced Raman spectroscopy.Spectrochim Acta A Mol Biomol Spectrosc. 2024 Mar 5;308:123786. doi: 10.1016/j.saa.2023.123786. Epub 2023 Dec 16. Spectrochim Acta A Mol Biomol Spectrosc. 2024. PMID: 38128327 Review.
-
Three-Dimensional Surface-Enhanced Raman Scattering Platforms: Large-Scale Plasmonic Hotspots for New Applications in Sensing, Microreaction, and Data Storage.Acc Chem Res. 2019 Jul 16;52(7):1844-1854. doi: 10.1021/acs.accounts.9b00163. Epub 2019 Jun 10. Acc Chem Res. 2019. PMID: 31180637
-
Self-Functionalized Superlattice Nanosensor Enables Glioblastoma Diagnosis Using Liquid Biopsy.ACS Nano. 2023 Oct 24;17(20):19832-19852. doi: 10.1021/acsnano.3c04118. Epub 2023 Oct 12. ACS Nano. 2023. PMID: 37824714
-
SERS as a Probe of Surface Chemistry Enabled by Surface-Accessible Plasmonic Nanomaterials.Acc Chem Res. 2023 Aug 1;56(15):2072-2083. doi: 10.1021/acs.accounts.3c00207. Epub 2023 Jul 12. Acc Chem Res. 2023. PMID: 37436068 Free PMC article.
Cited by
-
Exosomal Proteins and miRNAs as Mediators of Amyotrophic Lateral Sclerosis.Front Cell Dev Biol. 2021 Sep 10;9:718803. doi: 10.3389/fcell.2021.718803. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 34568332 Free PMC article. Review.
-
Solar-Powered CO2 Capture and Release Using Amine-Functionalized Plasmonic Titanium Nitride Nanoparticles.JACS Au. 2025 Jun 23;5(7):3027-3031. doi: 10.1021/jacsau.5c00489. eCollection 2025 Jul 28. JACS Au. 2025. PMID: 40747036 Free PMC article.
-
Multi-dimensional analysis of single particles with sequential dual-nanopipette sensors.Chem Sci. 2025 Jun 30;16(31):14170-14177. doi: 10.1039/d5sc02604k. eCollection 2025 Aug 6. Chem Sci. 2025. PMID: 40630633 Free PMC article.
-
Quantification of cyclic DNA polymerization with lanthanide coordination nanomaterials for liquid biopsy.Chem Sci. 2020 Mar 13;11(14):3745-3751. doi: 10.1039/c9sc06408g. Chem Sci. 2020. PMID: 34094063 Free PMC article.
-
Multiplex detection of ctDNA mutations in plasma of colorectal cancer patients by PCR/SERS assay.Nanotheranostics. 2020 Aug 25;4(4):224-232. doi: 10.7150/ntno.48905. eCollection 2020. Nanotheranostics. 2020. PMID: 32923312 Free PMC article.
References
-
- Brock G., Castellanos‐Rizaldos E., Hu L., Coticchia C., Skog J., Transl. Cancer Res. 2015, 4, 280.
-
- Otto A., J. Raman Spectrosc. 2005, 36, 497.
-
- Morton S. M., Jensen L., J. Am. Chem. Soc. 2009, 131, 4090. - PubMed
-
- Shan B., Pu Y., Chen Y., Liao M., Li M., Coord. Chem. Rev. 2018, 371, 11.
Publication types
LinkOut - more resources
Full Text Sources
Miscellaneous
