NBS1 interacts with HP1 to ensure genome integrity
- PMID: 31836699
- PMCID: PMC6911104
- DOI: 10.1038/s41419-019-2185-x
NBS1 interacts with HP1 to ensure genome integrity
Abstract
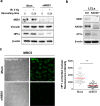
Heterochromatin Protein 1 (HP1) and the Mre11-Rad50-Nbs1 (MRN) complex are conserved factors that play crucial role in genome stability and integrity. Despite their involvement in overlapping cellular functions, ranging from chromatin organization, telomere maintenance to DNA replication and repair, a tight functional relationship between HP1 and the MRN complex has never been elucidated. Here we show that the Drosophila HP1a protein binds to the MRN complex through its chromoshadow domain (CSD). In addition, loss of any of the MRN members reduces HP1a levels indicating that the MRN complex acts as regulator of HP1a stability. Moreover, overexpression of HP1a in nbs (but not in rad50 or mre11) mutant cells drastically reduces DNA damage associated with the loss of Nbs suggesting that HP1a and Nbs work in concert to maintain chromosome integrity in flies. We have also found that human HP1α and NBS1 interact with each other and that, similarly to Drosophila, siRNA-mediated inhibition of NBS1 reduces HP1α levels in human cultured cells. Surprisingly, fibroblasts from Nijmegen Breakage Syndrome (NBS) patients, carrying the 657del5 hypomorphic mutation in NBS1 and expressing the p26 and p70 NBS1 fragments, accumulate HP1α indicating that, differently from NBS1 knockout cells, the presence of truncated NBS1 extends HP1α turnover and/or promotes its stability. Remarkably, an siRNA-mediated reduction of HP1α in NBS fibroblasts decreases the hypersensitivity to irradiation, a characteristic of the NBS syndrome. Overall, our data provide an unanticipated evidence of a close interaction between HP1 and NBS1 that is essential for genome stability and point up HP1α as a potential target to counteract chromosome instability in NBS patient cells.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures







Similar articles
-
The Drosophila Nbs protein functions in multiple pathways for the maintenance of genome stability.Genetics. 2006 Jul;173(3):1447-54. doi: 10.1534/genetics.106.058081. Epub 2006 Apr 30. Genetics. 2006. PMID: 16648644 Free PMC article.
-
The Drosophila Mre11/Rad50 complex is required to prevent both telomeric fusion and chromosome breakage.Curr Biol. 2004 Aug 10;14(15):1360-6. doi: 10.1016/j.cub.2004.07.019. Curr Biol. 2004. PMID: 15296753
-
Regulation of Mre11/Rad50 by Nbs1: effects on nucleotide-dependent DNA binding and association with ataxia-telangiectasia-like disorder mutant complexes.J Biol Chem. 2003 Nov 14;278(46):45171-81. doi: 10.1074/jbc.M308705200. Epub 2003 Sep 8. J Biol Chem. 2003. PMID: 12966088
-
Ataxia-telangiectasia-like disorder (ATLD)-its clinical presentation and molecular basis.DNA Repair (Amst). 2004 Aug-Sep;3(8-9):1219-25. doi: 10.1016/j.dnarep.2004.04.009. DNA Repair (Amst). 2004. PMID: 15279810 Review.
-
The importance of making ends meet: mutations in genes and altered expression of proteins of the MRN complex and cancer.Mutat Res. 2008 Sep-Oct;659(3):262-73. doi: 10.1016/j.mrrev.2008.05.005. Epub 2008 Jun 23. Mutat Res. 2008. PMID: 18606567 Review.
Cited by
-
IL-17A produced by POMC neurons regulates diet-induced obesity.iScience. 2024 Jun 12;27(7):110259. doi: 10.1016/j.isci.2024.110259. eCollection 2024 Jul 19. iScience. 2024. PMID: 39027371 Free PMC article.
-
A Tale of Ice and Fire: The Dual Role for 17β-Estradiol in Balancing DNA Damage and Genome Integrity.Cancers (Basel). 2021 Mar 30;13(7):1583. doi: 10.3390/cancers13071583. Cancers (Basel). 2021. PMID: 33808099 Free PMC article. Review.
-
Early differential responses elicited by BRAFV600E in adult mouse models.Cell Death Dis. 2022 Feb 10;13(2):142. doi: 10.1038/s41419-022-04597-z. Cell Death Dis. 2022. PMID: 35145078 Free PMC article.
-
Phosphopeptide interactions of the Nbs1 N-terminal FHA-BRCT1/2 domains.Sci Rep. 2021 Apr 27;11(1):9046. doi: 10.1038/s41598-021-88400-7. Sci Rep. 2021. PMID: 33907233 Free PMC article.
-
HP1γ Prevents Activation of the cGAS/STING Pathway by Preserving Nuclear Envelope and Genomic Integrity in Colon Adenocarcinoma Cells.Int J Mol Sci. 2023 Apr 16;24(8):7347. doi: 10.3390/ijms24087347. Int J Mol Sci. 2023. PMID: 37108510 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

