High frequency DNA rearrangement at qγ27 creates a novel allele for Quality Protein Maize breeding
- PMID: 31840105
- PMCID: PMC6904753
- DOI: 10.1038/s42003-019-0711-0
High frequency DNA rearrangement at qγ27 creates a novel allele for Quality Protein Maize breeding
Abstract
Copy number variation (CNV) is a major source of genetic variation and often contributes to phenotypic variation in maize. The duplication at the 27-kDa γ-zein locus (qγ27) is essential to convert soft endosperm into hard endosperm in quality protein maize (QPM). This duplication is unstable and generally produces CNV at this locus. We conducted genetic experiments designed to directly measure DNA rearrangement frequencies occurring in males and females of different genetic backgrounds. The average frequency with which the duplication rearranges to single copies is 1.27 × 10-3 and varies among different lines. A triplication of γ27 gene was screened and showed a better potential than the duplication for the future QPM breeding. Our results highlight a novel approach to directly determine the frequency of DNA rearrangements, in this case resulting in CNV at the qγ27 locus. Furthermore, this provides a highly effective way to test suitable parents in QPM breeding.
Keywords: Agricultural genetics; Genome duplication.
© The Author(s) 2019.
Conflict of interest statement
Competing interestsThe authors declare no competing interests.
Figures


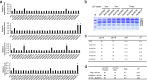
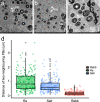


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

