Hybridization promotes asexual reproduction in Caenorhabditis nematodes
- PMID: 31841515
- PMCID: PMC6946170
- DOI: 10.1371/journal.pgen.1008520
Hybridization promotes asexual reproduction in Caenorhabditis nematodes
Abstract
Although most unicellular organisms reproduce asexually, most multicellular eukaryotes are obligately sexual. This implies that there are strong barriers that prevent the origin or maintenance of asexuality arising from an obligately sexual ancestor. By studying rare asexual animal species we can gain a better understanding of the circumstances that facilitate their evolution from a sexual ancestor. Of the known asexual animal species, many originated by hybridization between two ancestral sexual species. The balance hypothesis predicts that genetic incompatibilities between the divergent genomes in hybrids can modify meiosis and facilitate asexual reproduction, but there are few instances where this has been shown. Here we report that hybridizing two sexual Caenorhabditis nematode species (C. nouraguensis females and C. becei males) alters the normal inheritance of the maternal and paternal genomes during the formation of hybrid zygotes. Most offspring of this interspecies cross die during embryogenesis, exhibiting inheritance of a diploid C. nouraguensis maternal genome and incomplete inheritance of C. becei paternal DNA. However, a small fraction of offspring develop into viable adults that can be either fertile or sterile. Fertile offspring are produced asexually by sperm-dependent parthenogenesis (also called gynogenesis or pseudogamy); these progeny inherit a diploid maternal genome but fail to inherit a paternal genome. Sterile offspring are hybrids that inherit both a diploid maternal genome and a haploid paternal genome. Whole-genome sequencing of individual viable worms shows that diploid maternal inheritance in both fertile and sterile offspring results from an altered meiosis in C. nouraguensis oocytes and the inheritance of two randomly selected homologous chromatids. We hypothesize that hybrid incompatibility between C. nouraguensis and C. becei modifies maternal and paternal genome inheritance and indirectly induces gynogenetic reproduction. This system can be used to dissect the molecular mechanisms by which hybrid incompatibilities can facilitate the emergence of asexual reproduction.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




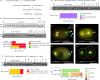

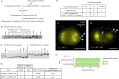

References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

