Critical Factors in Human Antizymes that Determine the Differential Binding, Inhibition, and Degradation of Human Ornithine Decarboxylase
- PMID: 31842334
- PMCID: PMC6995573
- DOI: 10.3390/biom9120864
Critical Factors in Human Antizymes that Determine the Differential Binding, Inhibition, and Degradation of Human Ornithine Decarboxylase
Abstract
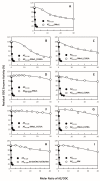
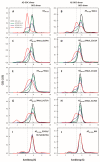
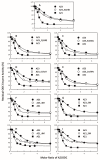
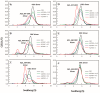
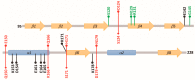
Antizyme (AZ) is a protein that negatively regulates ornithine decarboxylase (ODC). AZ achieves this inhibition by binding to ODC to produce AZ-ODC heterodimers, abolishing enzyme activity and targeting ODC for degradation by the 26S proteasome. In this study, we focused on the biomolecular interactions between the C-terminal domain of AZ (AZ95-228) and ODC to identify the functional elements of AZ that are essential for binding, inhibiting and degrading ODC, and we also identified the crucial factors governing the differential binding and inhibition ability of AZ isoforms toward ODC. Based on the ODC inhibition and AZ-ODC binding studies, we demonstrated that amino acid residues reside within the α1 helix, β5 and β6 strands, and connecting loop between β6 and α2 (residues 142-178), which is the posterior part of AZ95-228, play crucial roles in ODC binding and inhibition. We also identified the essential elements determining the ODC-degradative activity of AZ; amino acid residues within the anterior part of AZ95-228 (residues 120-145) play crucial roles in AZ-mediated ODC degradation. Finally, we identified the crucial factors that govern the differential binding and inhibition of AZ isoforms toward ODC. Mutagenesis studies of AZ1 and AZ3 and their binding and inhibition revealed that the divergence of amino acid residues 124, 150, 166, 171, and 179 results in the differential abilities of AZ1 and AZ3 in the binding and inhibition of ODC.
Keywords: AZ isoform; binding affinity; protein–protein interaction; ubiquitin-independent degradation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Critical factors governing the difference in antizyme-binding affinities between human ornithine decarboxylase and antizyme inhibitor.PLoS One. 2011 Apr 28;6(4):e19253. doi: 10.1371/journal.pone.0019253. PLoS One. 2011. PMID: 21552531 Free PMC article.
-
Structural basis of antizyme-mediated regulation of polyamine homeostasis.Proc Natl Acad Sci U S A. 2015 Sep 8;112(36):11229-34. doi: 10.1073/pnas.1508187112. Epub 2015 Aug 24. Proc Natl Acad Sci U S A. 2015. PMID: 26305948 Free PMC article.
-
A unified pathway for the degradation of ornithine decarboxylase in reticulocyte lysate requires interaction with the polyamine-induced protein, ornithine decarboxylase antizyme.Eur J Biochem. 1994 Dec 1;226(2):547-54. doi: 10.1111/j.1432-1033.1994.tb20079.x. Eur J Biochem. 1994. PMID: 8001569
-
Degradation of ornithine decarboxylase by the 26S proteasome.Biochem Biophys Res Commun. 2000 Jan 7;267(1):1-6. doi: 10.1006/bbrc.1999.1706. Biochem Biophys Res Commun. 2000. PMID: 10623564 Review.
-
The antizyme family for regulating polyamines.J Biol Chem. 2018 Nov 30;293(48):18730-18735. doi: 10.1074/jbc.TM118.003339. Epub 2018 Oct 24. J Biol Chem. 2018. PMID: 30355739 Free PMC article. Review.
Cited by
-
Protein degradation of antizyme depends on the N-terminal degrons.Protein Sci. 2024 Nov;33(11):e5199. doi: 10.1002/pro.5199. Protein Sci. 2024. PMID: 39473024
-
C-Methylated Spermidine Derivatives: Convenient Syntheses and Antizyme-Related Effects.Biomolecules. 2023 May 31;13(6):916. doi: 10.3390/biom13060916. Biomolecules. 2023. PMID: 37371496 Free PMC article.
-
The C-Terminus of the PSMA3 Proteasome Subunit Preferentially Traps Intrinsically Disordered Proteins for Degradation.Cells. 2022 Oct 14;11(20):3231. doi: 10.3390/cells11203231. Cells. 2022. PMID: 36291102 Free PMC article.
-
Structure simulation-based comparison of active site variations in fungal ornithine decarboxylases.Commun Integr Biol. 2025 Jan 31;18(1):2458872. doi: 10.1080/19420889.2025.2458872. eCollection 2025. Commun Integr Biol. 2025. PMID: 39906711 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

