SWR1 Chromatin Remodeling Complex: A Key Transcriptional Regulator in Plants
- PMID: 31842357
- PMCID: PMC6952815
- DOI: 10.3390/cells8121621
SWR1 Chromatin Remodeling Complex: A Key Transcriptional Regulator in Plants
Abstract
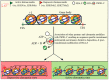
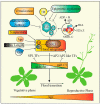
The nucleosome is the structural and fundamental unit of eukaryotic chromatin. The chromatin remodeling complexes change nucleosome composition, packaging and positioning to regulate DNA accessibility for cellular machinery. SWI2/SNF2-Related 1 Chromatin Remodeling Complex (SWR1-C) belongs to the INO80 chromatin remodeling family and mainly catalyzes the exchange of H2A-H2B with the H2A.Z-H2B dimer. The replacement of H2A.Z into nucleosomes affects nucleosome stability and chromatin structure. Incorporation of H2A.Z into the chromatin and its physiochemical properties play a key role in transcriptional regulation during developmental and environmental responses. In Arabidopsis, various studies have uncovered several pivotal roles of SWR1-C. Recently, notable progress has been achieved in understanding the role of SWR1-C in plant developmental and physiological processes such as DNA damage repair, stress tolerance, and flowering time. The present article introduces the SWR1-C and comprehensively reviews recent discoveries made in understanding the function of the SWR1 complex in plants.
Keywords: SWR1-C, H2A.Z; chromatin remodeling complex; flowering time regulation.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures
Similar articles
-
A plant-specific SWR1 chromatin-remodeling complex couples histone H2A.Z deposition with nucleosome sliding.EMBO J. 2020 Apr 1;39(7):e102008. doi: 10.15252/embj.2019102008. Epub 2020 Mar 2. EMBO J. 2020. PMID: 32115743 Free PMC article.
-
Roles of the INO80 and SWR1 Chromatin Remodeling Complexes in Plants.Int J Mol Sci. 2019 Sep 17;20(18):4591. doi: 10.3390/ijms20184591. Int J Mol Sci. 2019. PMID: 31533258 Free PMC article. Review.
-
Methyl-CpG-binding domain 9 (MBD9) is required for H2A.Z incorporation into chromatin at a subset of H2A.Z-enriched regions in the Arabidopsis genome.PLoS Genet. 2019 Aug 5;15(8):e1008326. doi: 10.1371/journal.pgen.1008326. eCollection 2019 Aug. PLoS Genet. 2019. PMID: 31381567 Free PMC article.
-
An insight into understanding the coupling between homologous recombination mediated DNA repair and chromatin remodeling mechanisms in plant genome: an update.Cell Cycle. 2021 Sep;20(18):1760-1784. doi: 10.1080/15384101.2021.1966584. Epub 2021 Aug 26. Cell Cycle. 2021. PMID: 34437813 Free PMC article. Review.
-
Regulation of MicroRNA-Mediated Developmental Changes by the SWR1 Chromatin Remodeling Complex.Plant Physiol. 2016 Jun;171(2):1128-43. doi: 10.1104/pp.16.00332. Epub 2016 Apr 18. Plant Physiol. 2016. PMID: 27208270 Free PMC article.
Cited by
-
New Insights into Cellular Functions of Nuclear Actin.Biology (Basel). 2021 Apr 7;10(4):304. doi: 10.3390/biology10040304. Biology (Basel). 2021. PMID: 33916969 Free PMC article. Review.
-
SDG2 regulates Arabidopsis inflorescence architecture through SWR1-ERECTA signaling pathway.iScience. 2021 Oct 12;24(11):103236. doi: 10.1016/j.isci.2021.103236. eCollection 2021 Nov 19. iScience. 2021. PMID: 34746701 Free PMC article.
-
OsChz1 acts as a histone chaperone in modulating chromatin organization and genome function in rice.Nat Commun. 2020 Nov 11;11(1):5717. doi: 10.1038/s41467-020-19586-z. Nat Commun. 2020. PMID: 33177521 Free PMC article.
-
Establishment and Maintenance of Heat-Stress Memory in Plants.Int J Mol Sci. 2024 Aug 18;25(16):8976. doi: 10.3390/ijms25168976. Int J Mol Sci. 2024. PMID: 39201662 Free PMC article. Review.
-
Chromatin Remodeling Complex SWR1 Regulates Root Development by Affecting the Accumulation of Reactive Oxygen Species (ROS).Plants (Basel). 2023 Feb 19;12(4):940. doi: 10.3390/plants12040940. Plants (Basel). 2023. PMID: 36840288 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

