Characterization of a live-attenuated HCMV-based vaccine platform
- PMID: 31848362
- PMCID: PMC6917771
- DOI: 10.1038/s41598-019-55508-w
Characterization of a live-attenuated HCMV-based vaccine platform
Abstract
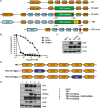
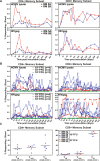
Vaccines based on cytomegalovirus (CMV) demonstrate protection in animal models of infectious disease and cancer. Vaccine efficacy is associated with the ability of CMV to elicit and indefinitely maintain high frequencies of circulating effector memory T cells (TEM) providing continuous, life-long anti-pathogen immune activity. To allow for the clinical testing of human CMV (HCMV)-based vaccines we constructed and characterized as a vector backbone the recombinant molecular clone TR3 representing a wildtype genome. We demonstrate that TR3 can be stably propagated in vitro and that, despite species incompatibility, recombinant TR3 vectors elicit high frequencies of TEM to inserted antigens in rhesus macaques (RM). Live-attenuated versions of TR3 were generated by deleting viral genes required to counteract intrinsic and innate immune responses. In addition, we eliminated subunits of a viral pentameric glycoprotein complex thus limiting cell tropism. We show in a humanized mouse model that such modified vectors were able to establish persistent infection but lost their ability to reactivate from latency. Nevertheless, attenuated TR3 vectors preserved the ability to elicit and maintain TEM to inserted antigens in RM. We further demonstrate that attenuated TR3 can be grown in approved cell lines upon elimination of an anti-viral host factor using small interfering RNA, thus obviating the need for a complementing cell line. In sum, we have established a versatile platform for the clinical development of live attenuated HCMV-vectored vaccines and immunotherapies.
Conflict of interest statement
O.H.S.U. and P.C., S.G.H., W.P., R.R., E.E.M., D.M., J.A.N., L.J.P. and K.F. have financial interests in Vir Biotechnology, Inc., a company that may have a commercial interest in the results of this research and technology. This potential conflict of interest has been reviewed and managed by O.H.S.U. All other authors declare no conflict of interest.
Figures





References
-
- Mendez, A. C., Rodriguez-Rojas, C. & Del Val, M. Vaccine vectors: the bright side of cytomegalovirus. Medical microbiology and immunology (2019). - PubMed
-
- Jarvis, M. A., Hansen, S. G., Nelson, J. A., Picker, L. J. & Früh, K. In Cytomegaloviruses: From Molecular Pathogenesis to Intervention Vol. 2 CYTOMEGALOVIRUSES: Molecular Biology and Immunology (ed Reddehase, M. J.) Ch. 21, (Caister Academic Press, 2013).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

