Expression Level of HIV-1 Vif Can Be Fluctuated by Natural Nucleotide Variations in the vif-Coding and Regulatory SA1D2prox Sequences of the Proviral Genome
- PMID: 31849897
- PMCID: PMC6893887
- DOI: 10.3389/fmicb.2019.02758
Expression Level of HIV-1 Vif Can Be Fluctuated by Natural Nucleotide Variations in the vif-Coding and Regulatory SA1D2prox Sequences of the Proviral Genome
Abstract
Vif is required for HIV-1 replication in natural target cells by counteracting host restriction factors, APOBEC3 (A3) proteins. We recently demonstrated that Vif expression level can be changed by naturally occurring single-nucleotide variations within SA1D2prox of the HIV-1 genome. We also found that levels for vif/vpr mRNAs are inversely correlated. While amino acid sequence per se is critical for functionality, Vif expression level modulated by signal sequences in its coding region is likely to be important as well. There are two splicing sites in the region involved in vpr expression. To reveal possible fluctuations of Vif-expression level, we examined SA1D2prox and vif gene by chimeric approaches using HIV-1 subtypes B and C with distinct anti-A3 activity. In this report, recombinant clones in subtype B backbone carrying chimeric sequences with respect to SA1D2prox/vif and those within the vif-coding region were generated. Of these, clones containing vif-coding sequence of subtype C, especially its 3' region, expressed vif/Vif at a decreased level but did at an increased level for vpr/Vpr. Clones with reduced vif/Vif level grew similarly or slightly better than a parental clone in weakly A3G-positive cells but more poorly in highly A3G-expressing cells. Three clones with this property were also tested for their A3-degrading activity. One of the clones appeared to have some defect in addition to the poor ability to express vif/Vif. Taken all together, our results show that natural variations in the SA1D2prox and vif-coding region can change the Vif-expression level and affect the HIV-1 replication potential.
Keywords: HIV-1; SA1D2prox; expression level; natural variation; subtype; vif.
Copyright © 2019 Doi, Koma, Adachi and Nomaguchi.
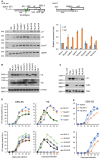
Figures

Similar articles
-
The Expression Level of HIV-1 Vif Is Optimized by Nucleotide Changes in the Genomic SA1D2prox Region during the Viral Adaptation Process.Viruses. 2021 Oct 15;13(10):2079. doi: 10.3390/v13102079. Viruses. 2021. PMID: 34696508 Free PMC article.
-
Production of HIV-1 vif mRNA Is Modulated by Natural Nucleotide Variations and SLSA1 RNA Structure in SA1D2prox Genomic Region.Front Microbiol. 2017 Dec 18;8:2542. doi: 10.3389/fmicb.2017.02542. eCollection 2017. Front Microbiol. 2017. PMID: 29326677 Free PMC article.
-
Natural Single-Nucleotide Variations in the HIV-1 Genomic SA1prox Region Can Alter Viral Replication Ability by Regulating Vif Expression Levels.J Virol. 2016 Apr 14;90(9):4563-4578. doi: 10.1128/JVI.02939-15. Print 2016 May. J Virol. 2016. PMID: 26912631 Free PMC article.
-
HIV-1 subtype variability in Vif derived from molecular clones affects APOBEC3G-mediated host restriction.Intervirology. 2013;56(4):258-64. doi: 10.1159/000348513. Epub 2013 May 8. Intervirology. 2013. PMID: 23689841
-
A naturally occurring Vif mutant (I107T) attenuates anti-APOBEC3G activity and HIV-1 replication.J Mol Biol. 2013 Aug 23;425(16):2840-52. doi: 10.1016/j.jmb.2013.05.015. Epub 2013 May 23. J Mol Biol. 2013. PMID: 23707381
Cited by
-
The Expression Level of HIV-1 Vif Is Optimized by Nucleotide Changes in the Genomic SA1D2prox Region during the Viral Adaptation Process.Viruses. 2021 Oct 15;13(10):2079. doi: 10.3390/v13102079. Viruses. 2021. PMID: 34696508 Free PMC article.
-
Interferon-Regulated Expression of Cellular Splicing Factors Modulates Multiple Levels of HIV-1 Gene Expression and Replication.Viruses. 2024 Jun 11;16(6):938. doi: 10.3390/v16060938. Viruses. 2024. PMID: 38932230 Free PMC article. Review.
-
Expression and Characterization of Two DNA Constructs Derived from HIV-1-vif in Escherichia coli and Mammalian Cells.Avicenna J Med Biotechnol. 2021 Jul-Sep;13(3):131-135. doi: 10.18502/ajmb.v13i3.6369. Avicenna J Med Biotechnol. 2021. PMID: 34484642 Free PMC article.
-
Commentary: Derivation of Simian Tropic HIV-1 Infectious Clone Reveals Virus Adaptation to a New Host.Front Cell Infect Microbiol. 2020 May 15;10:235. doi: 10.3389/fcimb.2020.00235. eCollection 2020. Front Cell Infect Microbiol. 2020. PMID: 32500043 Free PMC article. No abstract available.
References
-
- Akari H., Uchiyama T., Fukumori T., Iida S., Koyama A. H., Adachi A. (1999). Pseudotyping human immunodeficiency virus type 1 by vesicular stomatitis virus G protein does not reduce the cell-dependent requirement of vif for optimal infectivity: functional difference between Vif and Nef. J. Gen. Virol. 80, 2945–2949. 10.1099/0022-1317-80-11-2945, PMID: - DOI - PubMed
LinkOut - more resources
Full Text Sources

