Three New Orbivirus Species Isolated from Farmed White-Tailed Deer (Odocoileus virginianus) in the United States
- PMID: 31861885
- PMCID: PMC7019857
- DOI: 10.3390/v12010013
Three New Orbivirus Species Isolated from Farmed White-Tailed Deer (Odocoileus virginianus) in the United States
Abstract
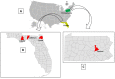
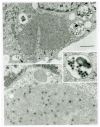
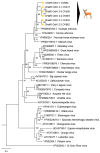
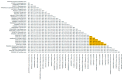
We report the detection and gene coding sequences of three novel Orbivirus species found in six dead farmed white-tailed deer in the United States. Phylogenetic analyses indicate that the new orbiviruses are genetically closely related to the Guangxi, Mobuck, Peruvian horse sickness, and Yunnan orbiviruses, which are thought to be solely borne by mosquitos. However, four of the six viruses analyzed in this work were found as co-infecting agents along with a known cervid pathogen, epizootic hemorrhagic disease virus-2 (EHDV-2), raising questions as to whether the new viruses are primary pathogens or secondary pathogens that exacerbate EHDV-2 infections. Moreover, EHDV-2 is known to be a Culicoides-borne virus, raising additional questions as to whether Culicoides species can also serve as vectors for the novel orbiviruses, if mosquitoes can vector EHDV-2, or whether the deer were infected through separate bites by the insects. Our findings expand knowledge of the possible viral pathogens of deer in the United States. Moreover, due to the close genetic relatedness of the three new orbiviruses to viruses that are primary pathogens of cattle and horses, our findings also underscore a crucial need for additional research on the potential role of the three new orbiviruses as pathogens of other animals.
Keywords: Odocoileus virginianus; deer farming; orbivirus; reovirus; white-tailed deer.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

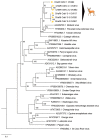



References
-
- Outlaw J.L., Anderson D.P., Earle M.L., Richardson J.W. Research Reoprt 17-3. Agricultural and Food Policy Center, Texas A&M University System; College Station, TX, USA: 2017. Economic impact of the Texas Deer Breeding and Hunting Operations.
-
- USDA . Health and Management Practices on U.S. Cervid Operations, 2014. USDA–APHIS–VS–CEAH–NAHMS; Fort Collins, CO, USA: 2016. No.698.0716.
-
- Seidl A.F., Koontz S.R. Potential Economic Impacts of Chronic Wasting Disease in Colorado. Hum. Dimens. Wildl. 2004;9:241–245. doi: 10.1080/10871200490480042. - DOI
-
- Adams K.P., Murphy B.P., Ross M.D. Captive White-Tailed Deer Industry—Current Status and Growing Threat. Wildl. Soc. Bull. 2016;40:14–19. doi: 10.1002/wsb.627. - DOI
-
- Campbell T.A., VerCauteren K. Diseases and Parasites. In: Hewitt D.G., editor. Biology and Management of White-Tailed Deer. CRC Press; Boca Raton, FL, USA: 2011. pp. 219–249.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical

